ADVERTISEMENTS:
This article throws light upon the top seven techniques used in genetic engineering. The seven techniques are: (1) Agarose Gel Electrophoresis (2) Isolation and Purification of Nucleic Acids (3) Isolation of Chromosomes (4) Nucleic Acid Blotting Techniques (5) DNA Sequencing (6) Alternative Methods of DNA Sequencing and (7) Chemical Synthesis of DNA.
Technique # 1. Agarose Gel Electrophoresis:
Electrophoresis refers to the movement of charged molecules in an electric field. The negatively charged molecules move towards the positive electrode while the positively charged molecules migrate towards the negative electrode.
Gel electrophoresis is a routinely used analytical technique for the separation/purification of specific DNA fragments. The gel is composed of either polyacrylamide or agarose. Polyacrylamide gel electrophoresis (PAGE) is used for the separation of smaller DNA fragments while agarose electrophoresis is convenient for the separation of DNA fragments ranging in size from 100 base pairs to 20 kb pairs. Gel electrophoresis can also be used for the separation of RNA molecules.
ADVERTISEMENTS:
Agarose is a polysaccharide derived from seaweeds. It forms a solid gel when dissolved in aqueous solution at concentrations between 0.5 and 2.0% (w/v). It may be noted here that the agarose used for electrophoresis is more purified form of agar when compared to that used for culture purposes. A diagrammatic view of the agarose gel electrophoresis unit is shown in Fig. 7.1.
The DNA samples are placed in the wells of the gel surface and the power supply is switched on. As the DNA is negatively charged, DNA fragments move through the gel towards the positive electrode. The rate of migration of DNA is dependent on the size and shape. In general, smaller linear fragments move faster than the larger ones. Hence, gel electrophoresis can be conveniently used for the separation of a mixture of DNA fragments, based on their size.
Agarose forms gels with pore sizes ranging from 100 to 300 nm in diameter. The actual pore size depends on the concentration of the agarose. The size of the pores determines the range of DNA fragments that can be separated on electrophoresis. For instance, a 0.3% agarose is used for the separation of DNA fragments between 5 and 50 kb, while a 5% agarose can separate 100-500 bp molecules.
ADVERTISEMENTS:
The migration of DNA fragments during the course of electrophoresis can be monitored by using dyes with known migration rates. These dyes are added to the DNA samples before loading. The bands of the DNA can be detected by soaking the gel in ethidium bromide solution.
When activated by ultraviolet radiation, DNA base pairs in association with ethidium bromide emit orange fluorescence. And in this way the DNA fragments separated in agarose electrophoresis can be identified (Fig. 7.2).
Pulsed-Field Gel Electrophoresis (PFGE):
The large sized DNA molecules (10 Mb) can be separated in agarose gels by using PFGE. This is made possible by periodically altering the direction of DNA migration by changing the orientation of electric field with respect to the gel. As the electric field orientation changes, the DNA molecules align themselves and migrate in new directions.
Limitation of PFGE:
The major limitation of pulsed-field gel electrophoresis is that the samples do not migrate in straight lines. This makes the further analysis of DNA rather difficult.
Contour-Clamped Homogeneous Electrical-Field Electrophoresis (CHEF):
CHEF is an improved method to apply alternating electrical fields to separate DNA molecules in electrophoresis. In this approach, the reorientation angle of electrical field is fixed at 120°C (or even at 90°) and electrophoresis carried out. By using CHEF, large DNA fragments (200-300 kb) can be routinely separated in a matter of hours.
Polyacrylamide Gel Electrophoresis (PAGE):
Polyacrylamide gel is composed of chains of acrylamide monomers cross-linked with methylenebisacrylamide units. The pore size of the gel is dependent on the total concentration of the monomers and the cross links. Polyacrylamide gel electrophoresis (PAGE) is used for the separation of single-stranded DNA molecules that differ in length by just one nucleotide.
Agarose gels cannot be used for this purpose. This is because polyacrylamide gels have smaller pore sizes than agarose gels and allow precise separation of DNA molecules from 10-1500 bp. Polyacrylamide gel electrophoresis is a wonderful technique for the fine separation of DNA molecules that differ from each other even by 1-3 nucleotides. PAGE is used in DNA sequencing and for the identification of amplified products of DNA by polymerase chain reaction (PCR).
Other Uses of Gel Electrophoresis:
ADVERTISEMENTS:
Besides the separation of DNA fragments, gel electrophoresis is also useful for understanding the molecular configuration of DNA molecules, and protein-nucleic acid interactions. The latter is based on the principle that binding of a protein to DNA usually results in the reduction of electrophoretic mobility.
The electrophoretic technique for protein- nucleic acid interaction involves the addition of a desired protein to double-stranded DNA fragments, separation of this complex and naked DNA by electrophoresis. The separated patterns can be visualized to understand the protein-nucleic acid interactions.
Technique # 2. Isolation and Purification of Nucleic Acids:
Almost all the experiments dealing with gene manipulations require pure forms of either DNA or RNA, or sometimes even both. Hence there is a need for the reliable isolation of nucleic acids from the cells. The purification of nucleic acids broadly involves three stages.
1. Breaking or opening of the cells to expose nucleic acids.
ADVERTISEMENTS:
2. Separation of nucleic acids from other cellular components.
3. Recovery of nucleic acids in a pure form.
Analytical procedures involving a few steps to several steps are in use for the purification of nucleic acids. In fact, commercial kits are readily available these days to enable purification of either DNA or RNA from different sources. The basic principles and procedures for nucleic acid purification are briefly described.
Purification of Cellular DNA:
The first step for DNA purification is to open the cells and release DNA. The method should be gentle to preserve the native DNA. Due to variability in cell structure, the approaches to break the cells are also different.
ADVERTISEMENTS:
Lysis of cells:
Bacterial cells:
The bacterial cells (e.g. E. coli) can be lysed by a combination of enzymatic and chemical treatments. The enzyme lysozyme and the chemical ethylenediamine tetra-acetate (EDTA) are used for this purpose. This is followed by the addition of a detergent such as sodium dodecyl sulfate (SDS).
Animal cells:
ADVERTISEMENTS:
Animal cells, particularly cultured animal cells, can be easily opened by direct treatment of cells with detergents (SDS).
Plant cells:
Plant cells with strong cell walls require harsh treatment to break open. The cells are frozen and then ground in a morter and pestle. This is an effective way of breaking the cellulose walls.
Methods to purify DNA:
There are two different approaches to purify DNA from the cellular extracts.
1. Purification of DNA by removing cellular components:
ADVERTISEMENTS:
This involves the degradation or complete removal of all the cellular components other than DNA. This approach is suitable if the cells do not contain large quantities of lipids and carbohydrates. The cellular extract is centrifuged at a low speed to remove the debris (e.g. pieces of cell wall) that forms a pellet at the bottom of the tube.
The supernatant is collected and treated with phenol to precipitate proteins at the interface between the organic and aqueous layers. The aqueous layer, containing the dissolved nucleic acids, is collected and treated with the enzyme ribonuclease (RNase). The RNA is degraded while the DNA remains intact. This DNA can be precipitated by adding ethanol and isolated after centrifugation, and suspended in an appropriate buffer.
2. Direct purification of DNA:
In this approach, the DNA itself is selectively removed from the cellular extract and isolated. There are two ways for direct purification of DNA. In one method, the addition of a detergent cetyltrimethyl ammonium (CTAB) results in the formation of an insoluble complex with nucleic acids. This complex, in the form of a precipitate is collected after centrifugation and suspended in a high-salt solution to release nucleic acids. By treatment with RNase, RNA is degraded. Pure DNA can be isolated by ethanol precipitation.
The second technique is based on the principle of tight binding between DNA and silica particles in the presence of a denaturing agent such as guanidinium thiocyanate. The isolation of DNA can be achieved by the direct addition of silica particles and guanidinium thiocyanate to the cellular extract, followed by centrifugation. Alternately, a column chromatography containing silica can be used, and through this the extract and guanidinium thiocyanate are passed. The DNA binds to the silica particles in the column which can be recovered.
Purification of Plasmid DNA:
Pure forms of plasmid DNA are often required in genetic engineering experiments. The prerequisite for a successful isolation of plasmid DNA is the appropriate and gentle lysis of the host cells, so that the plasmid DNA is released without too much contamination of chromosomal DNA.
ADVERTISEMENTS:
The high molecular weight host cell chromosomal DNA can be removed along with the cell debris by high-speed centrifugation. This results in the formation of a cleared lysate from which pure plasmid DNA can be isolated. Of the several methods in use to isolate plasmid DNA, two are briefly described.
Isopycnic centrifugation method:
The cleared lysate (referred above) is treated with a solution of cesium chloride (CsCI) containing ethidium bromide (EtBr). EtBr can bind with linear DNA molecules (chromosomal DNA fragments and open plasmid DNA) and not with circular plasmid DNA. The density of DNA-EtBr complex is much lower than the plasmid DNA. The circular plasmid DNA can be separated by isopycnic centrifugation in a CsCI-EtBr gradient (Fig. 7.3).
Birnboim and Doly method:
The technique developed by Birnboim and Doly (1979) is widely used for the purification of plasmid DNA. This method is based on the principle that there is a narrow range of pH (12.0-12.5) at which denaturation of linear DNA and not closed circular DNA occurs.
When the cleared lysate is subjected to carefully monitored alkaline pH (12.0 to 12.5), the plasmid DNA molecules remain in solution while all other DNA molecules get denatured and precipitate. This precipitate can be removed by centrifugation. The plasmid DNA in the supernatant can be concentrated by ethanol precipitation. Sometimes, further purification of plasmid DNA may be necessary which can be achieved by gel filtration.
Factors affecting plasmid DNA purification:
Several factors influence the purification of plasmid DNA. The most important ones are briefly described.
1. Plasmid copy number:
The number of plasmids present in the host cells at the time of harvest is important. The copy number in turn is influenced by growth medium, genotype of host cell and the stage of growth. In general, the higher the copy number, the better is the yield of plasmid DNA on purification.
2. Preparation of cleared lysate:
The methodology adopted and the care taken for the lysis of the host cells to prepare cleared lysate is important.
3. Host cells with end A gene:
The wild-type host cells possess endA gene that encodes the enzyme endonuclease I. The functions of this enzyme are not clearly known. It is however, observed that cell strains bearing endA mutations improve the stability and yield of plasmid DNA.
Purification of mRNA:
Among the RNAs, mRNA is frequently required in a pure form for genetic experiments. After the cells are disrupted on lysis by different techniques the cellular extract is deproteinised by treatment with phenol or phenol/ chloroform mixtures. On centrifugation, the nucleic acids get concentrated in the upper aqueous phase which may then be precipitated by using isopropanol or ethanol.
The purification of mRNA can be achieved by affinity chromatography using oligo (dT)-cellulose (Fig. 7.4). This is based on the principle that oligo (dT)-cellulose can specifically bind to the poly (A) tails of eukaryotic mRNA. Thus, by this approach, it is possible to isolate mRNA from DNA, rRNA and tRNA.
As the nucleic acid solution is passed through an affinity chromotographic column, the oligo (dT) binds to poly(A) tails of mRNA. By washing the column with high-salt buffer, DNA, rRNA and tRNA can be eluted, while the mRNA is tightly bound. This mRNA can be then eluted by washing with low-salt buffer. The mRNA is precipitated with ethanol and collected by centrifugation (Fig. 7.4).
Technique # 3. Isolation of Chromosomes:
Separation of large chromosomes of eukaryotes is not possible by conventional electrophoresis. The individual chromosomes of eukaryotes can be separated by fluorescence-activated cell sorting (FACS), also known as flow cytometry or flow karyotyping.
Fluorescence-Activated Cell Sorting:
To carry out FACS, the dividing cells (with condensed chromosomes) are carefully broken open, and a mixture of intact chromosomes is prepared. These chromosomes are then stained with a fluorescent dye. The quantity of the dye that binds to a chromosome depends on its size. Thus, larger chromosomes (with more DNA) bind more dye and fluoresce more brightly than the smaller ones.
The dye-mixed chromosomes are diluted and passed through a fine aperture that results in the formation of a stream of droplets. Each droplet contains a single chromosome. The fluorescence of the chromosomes is detected by a laser.
When the fluorescence indicates that the chromosome illuminated by the laser is the one desired, electrical charge is specifically applied to these droplets (and no others) which get charged. This results in the deflection of the droplets with the desired chromosome which can be separated from the rest and collected (Fig. 7.5). Uncharged droplets that do not contain the desired chromosome pass through a waste collection vessel.
Collection of Chromosomes with Identical Size:
The direct application of FACS (described above) is not suitable for the separation of chromosomes with identical sizes, e.g. chromosomes 21 and 22 in humans. Collection of such chromosomes can be achieved by use of special dyes (e.g. Hoechst 33258 and chromomycin A3) which bind to AT-rich DNA or GC-rich DNA. The rest of the procedure is the same that has been described. It is convenient to separate two or more chromosomes with identical sizes by different dyes. This is possible since no two chromosomes are likely to contain identical GC/AT contents.
Technique # 4. Nucleic Acid Blotting Techniques:
Blotting techniques are very widely used analytical tools for the specific identification of desired DNA or RNA fragments from thousands of molecules. Blotting refers to the process of immobilization of sample nucleic acids or solid support (nitrocellulose or nylon membranes). The blotted nucleic acids are then used as targets in the hybridization experiments for their specific detection. An outline of the nucleic acid blotting technique is depicted in Fig. 7.6.
Types of blotting techniques:
The most commonly used blotting techniques are listed below:
I. Southern blotting (for DNA)
II. Northern blotting (for RNA)
III. Dot blotting (DNA/RNA)
IV. Colony and plaque hybridization (cells from colony).
The Southern blotting is named after the scientist Ed Southern (1975) who developed it. The other names Northern blotting and Western blotting are laboratory jargons which are now accepted. Western blotting involves the transfer of protein blots and their identification by using specific antibodies.
A diagrammatic representation of a typical blotting apparatus is depicted in Fig. 7.7.
I. Southern Blotting:
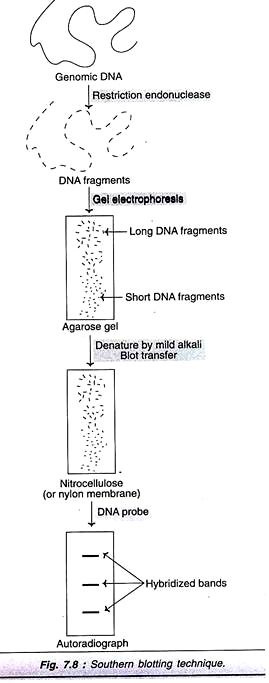
Southern blotting technique is the first nucleic acid blotting procedure developed in 1975 by Southern. It is depicted in Fig. 7.8, and briefly described.
The genomic DNA isolated from cells/tissues is digested with one or more restriction enzymes. This mixture is loaded into a well in an agarose or polyacrylamide gel and then subjected to electrophoresis. DNA, being negatively charged migrates towards the anode (positively charged electrode); the smaller DNA fragments move faster.
The separated DNA molecules are denatured by exposure to a mild alkali and transferred to nitrocellulose or nylon paper. This results in an exact replica of the pattern of DNA fragments on the gel. The DNA can be annealed to the paper on exposure to heat (80°C). The nitrocellulose or nylon paper is then exposed to labeled cDNA probes. These probes hybridize with complementary DNA molecules on the paper. The paper after thorough washing is exposed to X-ray film to develop autoradiograph. This reveals specific bands corresponding to the DNA fragments recognized by cDNA probe.
Factors affecting Southern blotting:
1. Stringent conditions (stringency control):
Stringency refers to the specificity with which a particular DNA target sequence is detected by a probe. Thus, with high stringent conditions (elevated temperature and low salt concentration) only completely complementary DNA sequences will bind and hybridize.
However, low stringent conditions will allow hybridization of partially matched sequences. Therefore, stringency control is very essential for specific detection of DNA molecules in Southern blotting. With good control of stringency, it is now possible to detect a DNA molecule just with a single base pair difference.
2. Membranes for blot transfer:
In the early years, nitrocellulose was used for immobilization of DNA molecules during blot transfer. The drawback with nitrocellulose is that it is fragile and has to be carefully handled. In recent years, most laboratories use nylon membranes as they possess high tensile strength, and better binding capacity for nucleic acids.
Applications of Southern blotting:
Southern blotting technique is extremely specific and sensitive, although it is a simple technique.
Some of the applications are listed:
a. It is an invaluable method in gene analysis.
b. Important for the confirmation of DNA cloning results.
c. Useful for mapping restriction sites around a single copy gene sequence.
d. Forensically applied to detect minute quantities of DNA (to identify parenthood, thieves, rapists etc.).
e. Highly useful for the determination of restriction fragment length polymorphism (RFLP) associated with pathological conditions.
f. DNA pieces from one species (e.g. human) can be used to detect DNA molecules from related species (e.g. chimpanzee, cow). This technique is referred to as Zoo-blotting.
II. Northern Blotting:
Northern blotting is the technique for the specific identification of RNA molecules. The procedure adopted is almost similar to that described for Southern blotting and is depicted in Fig. 7.9. RNA molecules are subjected to electrophoresis, followed by blot transfer, hybridization and autoradiography.
RNA molecules do not easily bind to nitrocellulose paper or nylon membranes. Blot- transfer of RNA molecules is carried out by using a chemically reactive paper prepared by diazotization of amino-benzyloxymethyl to create diazobenzyloxymethyl (DBM) paper. The RNA can covalently bind to DBM paper.
Some workers have later developed appropriate conditions to blot RNA bands on nitrocellulose paper, and modified nylon membranes. These are now widely used in RNA blotting. The use of DBM papers is almost discontinued. The blot-transferred RNA molecules hybridize with DNA probes which can be detected by autoradiography.
Northern blotting is theoretically, a good technique for determining the number of genes (through mRNA) present on a given DNA. But this is not really practicable since each gene may give rise to two or more RNA transcripts. Another drawback is the presence of exons and introns.
III. Dot-Blotting:
Dot-blotting is a modification of Southern and Northern blotting techniques described above. In this approach, the nucleic acids (DNA or RNA) are directly spotted onto the filters, and not subjected to electrophoresis. The hybridization procedure is the same as in original blotting techniques. Dot-blotting technique is particularly useful in obtaining quantitative data for the evaluation of gene expression.
IV. Western Blotting:
Western blotting involves the identification of proteins. It is to very useful to understand the nucleic acid functions, particularly during the course of gene manipulations. The technique of Western blotting involves the transfer of electrophoresed protein bands from polyacrylamide gel to nylon or nitrocellulose membrane. These proteins can be detected by specific protein-ligand interactions. Antibodies or lectins are commonly used for this purpose.
Autoradiography:
Autoradiography is the process of localization and recording of a radiolabel within a solid specimen, with the production of an image in a photographic emulsion. These emulsions are composed of silver halide crystals suspended in gelatin. When a p-particle or a ƴ-ray from a radiolabel passes through the emulsions, silver ions are converted to metallic silver atoms. This results in the development of a visible image which can be easily detected.
Direct autoradiography:
Direct autoradiography is ideally suited for the detection of weak to medium strength β-emitting radionuclides (3H, 14C, 35S). In this technique, the sample is placed in direct contact with the film. The radioactive emissions result in the development of black areas.
Indirect autoradiography:
For the detection of highly energetic β-particles (e.g. 32P, 125l), direct autoradiography is not suitable. This is because these emissions pass through and beyond the film, and a major part of the energy gets wasted. Indirect autoradiography is useful for the detection of highly energetic β-particles. In this technique, the β-particle energy is first converted to light by a scintillator, which then emits photons on exposure to photographic emulsion.
Applications of autoradiography:
As already described, autoradiography is closely associated with blotting techniques for the detection of DNA, RNA and proteins.
Colony and Plaque Blotting:
Colony and plaque blotting is a process of hybridization for the specific identification and purification of colonies (e.g. bacterial clones). This technique is depicted in Fig. 7.10, and briefly described hereunder.
The desired bacteria are grown as colonies on an agar plate. When a nitrocellulose filter paper is overlaid on the agar plate, the colonies get transferred. They are permanently fixed to the paper by heat. On treatment with alkali (NaOH), the cells lyse and the DNA gets denatured.
When these DNA prints are exposed to specific probes (radiolabel), hybridization occurs. The hybrid complex can be localized and detected by autoradiography. A similar strategy (described above for bacteria) can be used for the identification of phages and fragments of phage libraries.
Technique # 5. DNA Sequencing:
Determination of nucleotide sequence in a DNA molecule is the basic and fundamental requirement in biotechnology. DNA sequencing is important to understand the functions of genes, and basis of inherited disorders. Further, DNA cloning and gene manipulation invariably require knowledge of accurate nucleotide sequence.
Maxam and Gilbert Technique:
The first DNA sequencing technique, using chemical reagents, was developed by Maxam and Gilbert (1977). This method is briefly described below (Fig. 7.11).
A strand of source DNA is labeled at one end with 32P. The two strands of DNA are then separated. The labeled DNA is distributed into four samples (in separate tubes). Each sample is subjected to treatment with a chemical that specifically destroys one (G, C) or two bases (A + G, T + C) in the DNA. Thus, the DNA strands are partially digested in four samples at sites G, A + G, T + C and C. This results in the formation of a series of labeled fragments of varying lengths.
The actual length of the fragment depends on the site at which the base is destroyed from the labeled end. Thus for instance, if there are C residues at positions 4, 7, and 10 away from the labeled end, then the treatment of DNA that specifically destroys C will give lebeled pieces of length 3, 6 and 9 bases. The labeled DNA fragments obtained in the four tubes are subjected to electrophoresis side by side and they are detected by autoradiograph. The sequence of the bases in the DNA can be constructed from the bands on the electrophoresis.
Di-deoxynucleotide Method:
Currently, the preferred technique for determining nucleotide sequence in DNA is the one developed by Sanger (1980). This is an enzymatic procedure commonly referred to as the di-deoxynucleotide method or chain termination method (Note: Fredrick Sanger won Nobel Prize twice, once for determining the structure of protein, insulin; the second time for sequencing the nucleotides in an RNA virus).
A di-deoxynucleotide is a laboratory-made chemical molecule that lacks a hydroxyl group at both the 2′ and 3′ carbons of the sugar (Fig. 7.12). This is in contrast to the natural deoxyribonucleotide that possesses at 3′ hydroxyl group on the sugar.
Termination role of di-deoxynucleotide:
In the normal process of DNA replication, an incoming nucleoside triphosphate is attached by its 5′-phosphate group to the 3′-hydroxyl group of the last nucleotide of the growing chain when a di-deoxynucleotide is incorporated to the growing chain, no further replication occurs. This is because di-deoxynucleotide, lacking a 3′-hydroxyl group, cannot form a phosphodiester bond and thus the DNA synthesis terminates.
Sequencing method:
The process of sequencing DNA by di-deoxynucleotide method is briefly described. A single-stranded DNA to be sequenced is chosen as a template. It is attached to a primer (a short length of DNA oligonucleotide) complementary to a small section of the template. The 3′-hydroxyl group of the primer initiates the new DNA synthesis.
DNA synthesis is carried out in four reaction tubes. Each tube contains the primed DNA, Klenow subunit (the larger fragment of DNA polymerase of E. coli), four di-deoxyribonucleotides (ddATP, ddCTP, ddGTP or ddTTP). It is necessary to radiolabel (with 32P) the primer or one of the deoxyribonucleotides.
As the new DNA synthesis is completed, each one of the tubes contains fragments of DNA of varying length bound to primer. Let us consider the first reaction tube with di-deoxyadenosine (ddATP). In this tube, DNA synthesis terminates whenever the growing chain incorporated ddA (complementary to dT on the template strand). Therefore, this tube will contain a series of different length DNA fragments, each ending with ddA. In a smilar fashion, for the other 3 reaction tubes, DNA synthesis stops as the respective di-deoxynucleotides are incorporated.
The synthesis of new DNA fragments in the four tubes is depicted in Fig. 7.13.
The DNA pieces are denatured to yield free strands with radiolabel. The samples from each tube are separated by polyacrylamide gel electrophoresis. This separation technique resolves DNA pieces, different in size even by a single nucleotide. The shortest DNA will be the fastest moving on the electrophoresis.
The sequence of bases in a DNA fragment is determined by identifying the electrophoretic (radiolabeled) bands by autoradiography. In the Fig. 7.14, the sequence of the newly synthesized DNA fragment that is complementary to the original DNA piece is shown.
It is conventional to read the bands from bottom to top in 5′ to 3′ direction. By noting the order of the bands first C, second G, third T and so on, the sequence of the DNA can be determined accurately. As many as 350 base sequences of a DNA fragment can be clearly identified by using autoradiographs.
Modifications of di-deoxynucleotide method:
Replacement of 32P-radiolabel by 33P or 35S improves the sharpness of auto-radiographic images. DNA polymerase of the thermophilic bacterium, Thermus aquaticus (in place of Klenow fragment of E. coli DNA polymerase I) or a modified form of phage T7 DNA polymerase (sequenase) improves the technique.
Limitations of di-deoxynucleotide method:
There are mainly two limitations in this techniques. The need for a single-stranded DNA template and the use of a primer to an unknown sequences. These problems can be overcome by using bacteriophage M13.
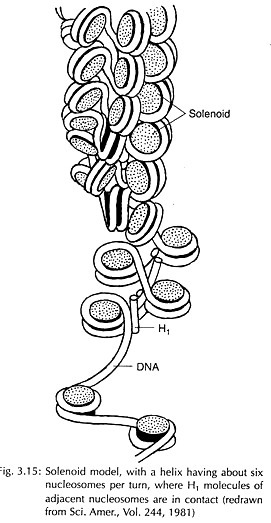
Bacteriophage M13 as A Cloning and DNA Sequencing Vector:
The life cycle of phage M13 is depicted in Fig. 7.15. This virus contains a single-stranded circular DNA. When it infects E. coli, the double- stranded DNA is synthesized which is a replicative form (RF). RF DNA replicates until 50-100 copies are produced. Now the mode of replication is changed to synthesize single-stranded DNA (ssDNA) molecules.
Each of the ssDNA of phage M13 is coated with a protein. The phage particles then freely pass through the membrane to be released out. Thus, the E. coli cells infected with M13 can continuously secrete infective particles, without undergoing lysis.
Bacteriophage M13 is employed as a cloning and DNA sequence determining vector. This is possible since the double-stranded replicative form can be isolated and used as a plasmid, while the single-stranded phage DNA can be employed as a template for DNA sequencing. Usually, a target DNA with less than 500 bp can be cloned and sequenced in phage M13.
This double-stranded RF forms of phage M13 are isolated and the DNA to be cloned is inserted. This phage which is comparable to a plasmid is returned to E. coli. (The uptake of phages with inserts can be screened by using lac Z marker).
As the phage undergoes its life cycle (Fig. 7.15), single-stranded DNA molecules of the inserted DNA are recovered. Sequence determination of the single-stranded DNA can be done by di-deoxynucleotide method .The primer used is complementary to a part of phage M13 DNA which is close to the point of insertion. Therefore, a single primer can be used for different DNA molecules inserted. For determining the sequence of a larger DNA (≤ 2,000 bp), it is necessary to clone different pieces of the DNA. From the sequences of these fragments (frequently overlapping), the final sequence can be built.
DNA Sequencing by Primer Walking:
Primer walking or primer extension technique is employed for sequence determination of long pieces of DNA (≥ 5,000 bp). In this method, plasmid cloned DNA containing target DNA is annealed with a primer (synthetic oligonucleotide). The primer binds with the DNA close to the inserted DNA. Now the target DNA (250-350 nucleotides) is sequenced, most frequently by di-deoxynucleotide method (described already).
From the base sequence of a fragment of the target DNA (determined in the first round), the second primer is chosen. It is an oligonucleotide to bind to a region approximately 300 nucleotides away from first primer binding. The next fragment of target DNA can be sequenced by another 250-300 bases (second round).
This sequence will be useful to choose the third primer and determine the next 250-350 bases (third round). In this fashion, the primer walking is continued until the complete target DNA is sequenced. Three rounds of primer walking technique for DNA sequencing are depicted in Fig. 7.16.
Chromosome Walking In DNA Sequencing:
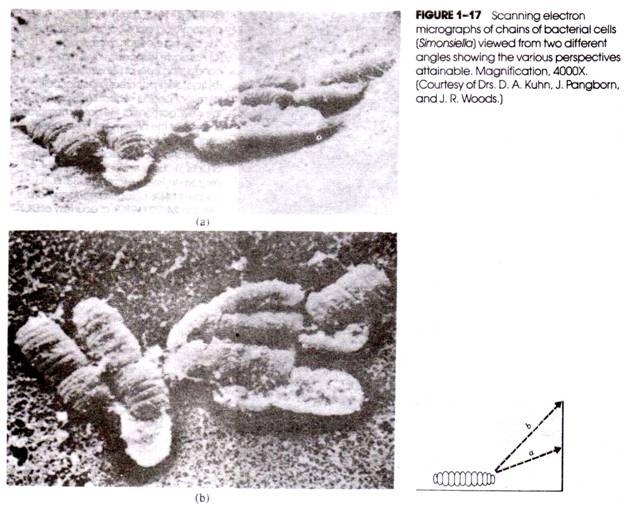
Chromosome walking is a DNA base sequencing method in which the chromosome is analysed by extending one tip to reach the other. The technique basically involves systematically moving along the chromosome from a known location to an unknown location, and thus determining the DNA sequence. The method adopted in chromosome walking is depicted in Fig. 7.17, and briefly described below.
A DNA fragment with approximately 40,000 base pairs can be sequenced. To start with a marker is used to identify the appropriate gene probe (from the DNA probe library). This gene probe must have a section of the base pair identical to the base pairs of the marker. This initial probe will hybridize only with the clones containing fragment A.
This fragment A can be isolated, cloned and used as a probe to detect fragment B. This procedure of cloning and probing with fragments is repeated again and again until fragment D hybridizes with fragment E. As is evident from the above description, chromosome walking uses probes derived from the ends of overlapping clones to facilitate a walk along with the DNA sequence. In the process of this walk, the sequence of the desired target gene can be identified.
Chromosome jumping:
An improved strategy of chromosome walking is chromosome jumping. Many regions of DNA that are difficult to be cloned by walking can be jumped. The procedure involves the circularization of large genomic fragments generated by the digestion of endonucleases. This is followed by cloning of the region lowering the closure of the fragment. By this approach, the DNA sequences located at far off places can be brought together and cloned. A chromosome jumping can be constructed in this manner and used for long distance chromosome walks.
Applications of chromosome walking and jumping:
The gene responsible for the disease cystic fibrosis (CF) has been identified by chromosome walking and jumping. The gene of protein responsible for CF is called cystic fibrosis trans membrane conductance regulator (CFTR) and is located on the long arm of chromosome 7.
Automated DNA Sequencing:
DNA sequencing in the recent years is carried out by an automated DNA sequencer. In this technique, fluorescent tags are attached to chain- terminating nucleotides (di-deoxynucleotides). This tag gets incorporated into the DNA molecules, while terminating new strand synthesis.
Four different fluorescent dyes are used to identify chain- terminating reactions in a sequencing gel. The DNA bands are separated by electrophoresis and detected by their fluorescence. Recently, four dyes that exhibit strong absorption in laser are in use for automated sequencing.
Advantages of automated sequencing:
It is a rapid and accurate technique. Automated DNA sequencer can accurately sequence up to 100,00 nucleotides per day. The cost works out to be not more than $0.2 per nucleotide. Automated DNA sequencing has been successfully used in the human genome project.
Technique # 6. Alternative Methods of DNA Sequencing:
Some groups of research workers have developed alternate methods to Sanger method for sequencing of DNA. Unfortunately, despite the initial excitement, most of these methods have disappeared from the scene. There are at least two methods with some promise for DNA sequencing- pyro-sequencing, and gene chips (microarrays).
Pyro-sequencing:
Pyro-sequencing is a DNA sequencing method that is based on the principle of determining which one of the four bases (A, G, C, T) is incorporated at each step while a DNA template is copied. In this technique, the template copies in a straight forward manner without added dideoxy- nucleotides (ddNTPs).
As the new strand is synthesized, the order in which the deoxynucleotides (dNTPs) are incorporated is detected, and the sequence can be read as the reaction proceeds (Fig. 7.18). The identification of base addition becomes possible since the addition of a nucleotide is accompanied by the release of a molecule of pyrophosphate. This can be detected by chemiluminescence technique.
In the procedure of pyro-sequencing, each dNTP is added individually (not all four together), along with a nucleotides enzyme. This enzyme degrades the dNTP if it is not incorporated into the newly synthesized DNA strand. Once the appropriate nucleotide is incorporated into the new DNA strand, a molecule of pyrophosphate is released.
This can be converted by the enzyme sulfurylase into a flash of light (chemiluminescence). By the addition of each dNTP separately, one after another, the order of the nucleotides added to the growing DNA strand can be followed, and the sequence can be identified.
Detection of the molecule pyrophosphate is the basis of DNA sequencing, hence the name pyro sequencing. Since pyro sequencing does not require electrophoresis or any other DNA fragment separation technique, it is more rapid than chain termination sequencing.
The major limitation of pyro sequencing is that it is suitable to detect the sequence of about 200 nucleotides. This is much less than the Sanger method. Many improvements are being made in pyro sequencing so that much longer DNA molecules can be sequenced. In fact, an automated system for pyro sequencing is also available now.
DNA Chips (Microarrays):
DNA chips or DNA microarrays are recent developments for DNA sequencing as result of advances made in automation and miniarization. A large number of DNA probes, each one with different sequence, are immobilized at defined positions on the solid surface, made up of either nylon or glass. The probes can be short DNA molecules such as cDNAs or synthetic oligonucleotides. For the preparation of high density arrays, oligonucleotides are synthesized in situ on the surface of glass or silicon. This results in an oligonucleotide chip rather than a DNA chip.
Technique of DNA sequencing:
A DNA chip carrying an array of different oligonucleotides can be used for DNA sequencing. For this purpose, a fluorescently labeled DNA test molecule, whose sequence is to be determined, is applied to the chip. Hybridization occurs between the complementary sequences of the test DNA molecule and oligonucleotides of the chip.
The positions of these hybridizing oligonucleotides can be determined by confocal microscopy. Each hybridizing oligonucleotide represents an 8-nucleotide sequence that is present in the DNA probe. The sequence of the test DNA molecule can be deduced from the overlaps between the sequences of the hybridizing oligonucleotides (Fig. 7.19).
Applications of DNA chips:
There have been many successes with this relatively new technology of DNA chips. Some of them are listed.
a. Identification of genes responsible for the development of nervous systems.
b. Detection of genes responsible for inflammatory diseases.
c. Construction of microarrays for every gene in the genome of E. coli, and almost all the genes of the yeast Saccharomyces cerevisiae.
d. Expression of several genes in prokaryotes has been identified.
e. Detection and screening of single nucleotide polymorphisms (SNPs).
f. Rapid detection of microorganisms for environmental monitoring.
The future of DNA chips:
The major limitation of DNA chips at present is the unavailability of complete genome arrays for higher eukaryotes, including humans. It is expected that within the next few years such DNA chips will be available. This will help the biotechnologists to capture the functional snapshots of the genome in action for higher organisms.
Technique # 7. Chemical Synthesis of DNA:
Advances in the laboratory techniques have made it possible to chemically synthesize DNA in a short period. Thus, oligonucleotides of about 100 bases can be produced in about 10 hours. Laboratory synthesis of DNA (recently by use of DNA synthesizers or gene machines) with specific sequence of nucleotides, rapidly and inexpensively, has significantly contributed to cloning. Chemical synthesis of DNA is based on the ability to protect the reactive -5′ and -3′ ends by blocking (protecting) them.
The Phosphoramidite Method:
The phosphoramidite method is the technique of choice, currently in use, for the synthesis of DNA. The synthesis is carried out on a solid phase system i.e. by attaching the growing DNA strand to a solid support, in a reaction vessel. The procedure involves the following stages.
1. Nucleoside attachment to solid support:
The initial nucleoside (base + sugar) is attached at 3′ end to an inert solid support, usually to glass beads with uniform pores, referred to as controlled pore glass (CPG) beads.
2. Preparation of phosphoramidites:
For each one of the four bases (A, G, C and T), phosphoramidites can be synthesized. This is achieved by attaching dimethoxytrityl (DMT) group to the 5′-hydroxyl group of deoxyribose and a diisopropylamine group to the 3′-phosphite group of the nucleoside. A methyl residue protects the 3′- phosphite group. The general structure of phosphoramidite is depicted in Fig. 7.20.
3. Coupling:
A nucleotide precursor (i.e. a phosphoramidite) with its 3′-phosphorus couples with 5′-hydroxyl of the initial nucleoside bound to a CPG bead.
4. Oxidation and de-protection:
The unstable trivalent phosphate is oxidized to pentavalent stable phosphate. This is followed by the removal of DMT that protects (de-protection) 5-hydroxyl group. For convenience, oxidation and de-protection are considered together in Fig. 7.21 also, they are actually two independent reactions. Another nucleotide precursor is now added and the reactions described in steps 3 and 4 are repeated.
Coupling, oxidation and de-protection are the three steps in the cyclic reaction for the DNA synthesis by phosphoramidite method. The cycles are repeated again and again to synthesise the desired DNA. At the end, the completed oligonucleotide is removed from the glass support and the groups protecting the bases and phosphates are cleaved.
Applications of synthesized oligonucleotides:
Chemically synthesized oligonucleotides have a number of uses in biotechnology.
1. The linkers and adaptors are the oligonucleotides commonly employed in the preparation of recombinant DNA
2. Chemically synthesized DNAs with known sequences can be used for the synthesis of proteins/ polypeptides.
3. DNA probes, particularly the single-stranded oligonucleotide probes, can be synthesized in the laboratory. This is based on the codon sequence of mRNA which in turn is dependent on the amino acid sequence.
4. Single-stranded oligonucleotides are used as primers in polymerase chain reaction (PCR).
5. For in vitro mutagenesis, single-stranded oligonucleotides are employed.
Synthesis of Genes:
Gene is a double-stranded DNA; two single- stranded complementary oligonucleotides can be synthesized, separately, and on annealing they form double strands. This can be conveniently achieved for smaller genes (60-90 bp). However, the synthesis of longer genes (> 300 bp) is associated with some practical difficulties.
This is mainly due to the fact that the coupling efficiency for the chemical synthesis of DNA is never 100%. To overcome this problem, small DNA fragments are synthesized and assembled (Fig. 7.22). Sealing of the nicks is achieved by use of T4 DNA ligase.
Another way of synthesizing genes is to produce overlapping oligonucleotides which on annealing contain large gaps. These gaps can be filled by enzymatic synthesis of DNA by DNA polymerase—while synthesizing genes in the laboratory, utmost care should be taken to see that the nucleotides are in the correct sequence (this can be checked by DNA sequence technique). An Indian born and American settled scientist Har Gobind Khorana and his associates were the first to synthesize an entire tRNA gene for yeast alanyl tRNA in 1972.