ADVERTISEMENTS:
Let us make an in-depth study of the RNA processing. After reading this article you will learn about: 1. Processing of Eukaryotic mRNA 2. RNA Splicing and Mechanisms of Splicing and 3. Processing of tRNA.
Almost all types of RNA molecules undergo post synthesis transformation which is called RNA processing. Prokaryotic mRNA is generally not processed. In prokaryotes 5′-end of prokaryotic mRNA starts translation while the 3′-end is still under synthesis. Eukaryotic mRNA undergoes maximum processing. Both prokaryotic and eukaryotic tRNAs and rRNAs undergo processing. Except prokaryotic mRNA, all other kinds of RNA are processed immediately after synthesis.
Processing of Eukaryotic mRNA:
Newly synthesized mRNA is called primary transcription or precursor mRNA. It is quite different from the mRNA that takes part in protein synthesis. Large-scale changes take place in precursor mRNA. These changes are called processing of mRNA. Both 5′-end 3′-end of mRNA are modified. Non-coding regions are removed by splicing. The changes lead to the formation of mature mRNA which takes part in protein synthesis.
5′-end Capping:
ADVERTISEMENTS:
At 5′-end of mRNA a cap of 7-methylguanosine (m7G or 7MeG) is added immediately after transcription or during transcription. This is most common and is called a cap. This cap is added in reverse orientation of 5′ → 5′ instead of normal 5′ → 3′ orientation giving a 5-5′ triphosphate bridge. A methyl group is added to the 7th position of terminal guanine. This reaction is catalysed by an enzyme guanylyl transferase. The cap provides stability to mRNA as the exonuclease is unable to act on it.
3′-Cleavage and Polyadenylation:
At the 3′-end of mRNA a tail consisting of sequence of adenine (adenylic acid) is added immediately after transcription. This tail is called poly (A) tail. It consists of 80-200 adenine nucleotides. This addition is catalysed by an enzyme poly (A) polymerase or PAP. The tail provides stability to mRNA.
RNA Splicing and Mechanisms of Splicing:
Till recently it was believed that coding sequences of DNA and amino acids of polypeptide chain are collinear. Recently it has been discovered that coding sequences of most of the eukaryotic genes is split into stretches of codons interrupted by stretches of non-coding sequences.
ADVERTISEMENTS:
The coding sequences of DNA are called exons. In between exons, there are intervening non-coding sequences called introns. This type of genes are called split genes or interrupted genes. The terms exons and introns were given by Gilbert. Introns are usually much larger than the exons. Moreover, the introns constitute a large portion of the genome. During processing of mRNA, the introns are excised out and removed by a process called RNA splicing. Translation takes place after the splicing is completed.
Processing of tRNA:
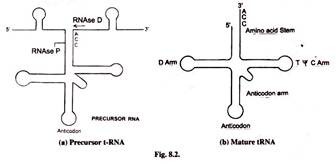
tRNA undergoes extensive processing. The mature tRNAs consists of 80-90 nucleotides. But the precursor tRNA is much longer. For example tRNATyr which is a tyrosine carrying tRNA contains 350 nucleotides. Processing discards useless sequences. This is done by enzymes RNase D, RNase E, RNase F and RNase P. Nucleotides are removed from both 5′ and 3′ ends.
Endonucleases also remove many sequences. Cleaving is done after the primary transcript has folded and formed characteristic stems and loop structure by extensive complementary base pairing. RNase P is a ribozyme.
The 5′-CCA-3′ sequence at 3′-end of mature tRNA is added by the enzyme tRNA nucleotidyl transferse. This generates the mature 3′-end the tRNA.
Several unusual bases are formed by the modification of normal existing bases A, G, C and U by the enzymatic action. These modified bases are pseudouridine (Ψ), 2- isopentenyladenosine (2 ip A), 2-O-methylguanosine (2m G), 4-thiouridine (4 t μ), Ribothymidine, dihyrouridine and inosine.
rRNA Processing:
In rRNA processing in both prokaryotes and eukaryotes, the primary transcript undergoes some changes. Some nucleotides are removed by exonucleases and endonucleases. New nucleotides are added both at 5′-end and 3′-end. Certain nucleotides are modified.
rRNA Processing in Prokaryotes:
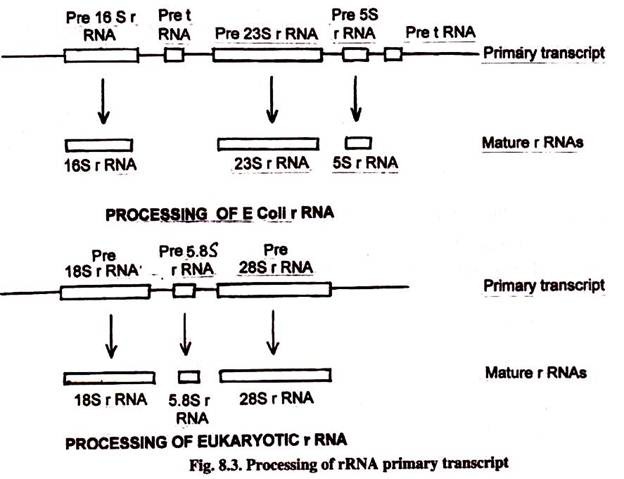
In E. coli there are several different operons in rRNA. Each operon contains one copy of 5S, 16S and 23S rRNA sequences. In addition, several coding sequences of tRNA molecules are also present in rRNA operons. These primary transcriptions are processed to give both rRNA and tRNA molecules. All these molecules are cleaved from a continuous transcript of more than 5000 nucleotides.
rRNA Processing in Eukaryotes:
The precursor rRNA in eukaryotes contain one copy of 18S coding region, one copy of 5.8S and one copy of 28S coding region. This rRNA precursor is transcribed in the nucleolus by RNA polymerase I. The eukaryotic 5S rRNA is transcribed by RNA polymerase III from an unlinked gene.
ADVERTISEMENTS:
In both prokaryotes and eukaryotes rRNA molecules form secondary structures of numerous double stranded stems by complementary base pairing and single stranded loops.
Small nucleolar RNA (snoRNA) is required for the processing of rRNA molecules in eukaryotes. The snoRNAs are present in the nucleolus where rRNA is processed. Ribosomes are also assembled in nucleolus.
The RNA molecules in cells are present in the form of complexes formed with proteins. Specific proteins bind to specific RNAs. The RNA-protein complexes are called ribonuclear proteins or RNPs. Ribosomes are the largest RNPs formed by rRNA molecules forming complexes with specific ribosomal proteins. In E. coli ribosomes account for 25% of the dry weight of the cell. It consists of 10% of total proteins and 80% of total RNA.
ADVERTISEMENTS:
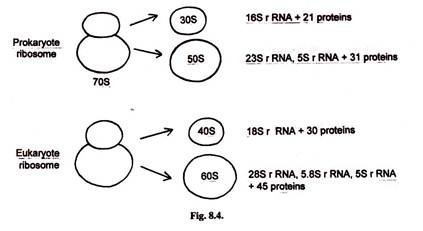
In prokaryotes 70S ribosome has 30S and 50S subunits. The 30S subunit has one copy of 16S rRNA and 21 different proteins. The 50S subunit has 23S rRNA, 5S rRNA and 31 different proteins.
In eukaryotes 80S ribosome has 40S and 60S subunits. The 40S subunit has 18S rRNA and 30 different proteins. The 60S subunit has 28S rRNA, 5.8S rRNA, 5.8S rRNA and 45 different proteins.
Summary:
Except prokaryotic mRNA all other kinds of RNAs are processed immediately after synthesis. Eukaryotic mRNA undergoes maximum processing. Both prokaryotic arid eukaryotic tRNAs and rRNAs undergo extensive processing. Processing of eukaryotic mRNA: Newly synthesized mRNA is called pre-mRNA. It is quite different from the mRNA that takes part in protein synthesis. At 5′ end of mRNA, a cap of 7-methylguanosine (m7G or 7 Me G) is added. At 3′-end a tail consisting of 80-200 adenine nucleotides called Poly(A) tail is added.
ADVERTISEMENTS:
Non-coding regions or introns are removed by splicing and coding regions or exons are joined together.
Processing of tRNA:
Pre-tRNA consists of upto 350 nucleotides. Processing discards most of useless nucleotides. Mature tRNA consists of about 70 nucleotides. At the 3′ end CCA sequence is added. Several unusual bases are found by the modification of normal bases A, G, C and U.
rRNA Processing:
In both prokaryotes and eukaryoties rRNA molecules form secondary structures of numerous double stranded stems by complementary base pairing and single stranded loops. Many proteins bind to these rRNA molecules to form ribonucleoproteins (RNP) in the ribosomes. There are several small nuclear RNA molecules called small nuclear RNAs (snRNA).