ADVERTISEMENTS:
Read this article to learn about the stages, primer design, types, sensitivity, factors affecting, applications and variations of polymerase chain reaction.
PCR has been one of the most important techniques developed in recent years. The reason behind is its simplicity of the reaction and relative case of the practical manipulation steps.
The PCR is used to amplify a precise fragment of DNA from a complex mixture of starting material usually termed as template DNA.
ADVERTISEMENTS:
The reaction requires some information about sequence of flanking fragment of DNA to be amplified. From this information two oligonucleotide primers can be synthesized each complimentary to the stretch of DNA to the 3′-side of the target DNA, one for each of the two DNA strands. This technique in many cases has replaced traditional DNA cloning methods, since it fulfills the same function of producing large amount of DNA from small amount of starting materials.
Stages in PCR:
It has three definite sets of times and temperature, termed as
Step I:
ADVERTISEMENTS:
Denaturation
Step II:
Annealing and
Step III:
Extension.
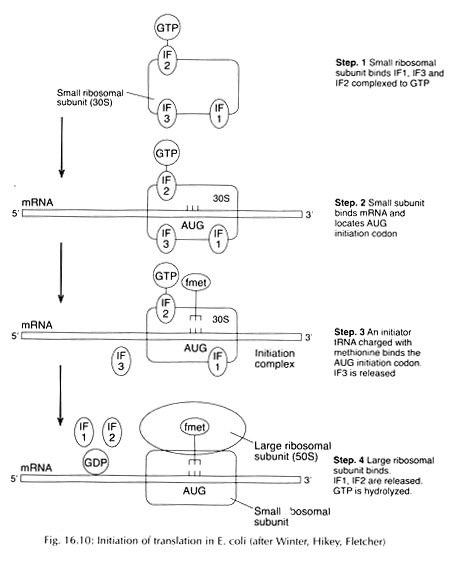
Each of the three steps are repeated 30-40 times or cycles. In first cycle the double stranded template DNA strand is first denatured by heating the reaction to above 90°C so that the region to be specifically amplified can be made accessible. The temperature is then cooled to between 40-60°C. The precise temperature is critical and each PCR system has to be defined and optimized. Reactions that are not optimized may give rise to other products along with amplified target sequence or even may not produce amplified product.
The second annealing step allows the hybridization of the two oligonucleotide primers, present in excess to bind to their complimentary sites which flank the DNA. The annealed oligonucleotide acts as a primer since they provide free 3′-end hydroxyl group to DNA polymerase. The third step, DNA synthesis or extension is carried out by a thermo stable DNA polymerase, most commonly Taq DNA polymerase.
DNA synthesis proceeds from both primers until new strands have been extended along and beyond the target DNA to be amplified. It is important to note that since the new strand extends beyond the target DNA, they will contain a region next to their 3′-end that is complimentary to the other primer. Thus in another round not only parental target strand but also newly synthesized strands also act as template DNA.
ADVERTISEMENTS:
One problem with the early PCR reaction was the temperature needed to denature the DNA also denatures the DNA polymerase. Then thermo stable DNA polymerase called Taq polymerase from thermophilic bacterium Thermus aquaticus with optimum temperature of 72°C and can also survive after prolonged exposure to 96°C provided the means to automate the reaction.
Primer Design in PCR:
The principle features o ideal primers are that these not only have to be complementary to the flanking sequence of the target DNA but must not be self-complementary otherwise they will form primer dimmers. Both primers have to be matched in their G+C content and should have similar annealing temperature.
It is also possible to design primer with additional sequences at their 5′-ends, such as restriction endonuclease target site or promoter sequences. However, these require alteration in annealing temperature to compensate for areas of non-homology.
Types of PCR:
The PCR may be used to amplify the templates from variety of sources. Unlike many manipulation methods used in current molecular biology the PCR technique is sensitive enough to require very little template preparation.
Extraction from many cells require just single boiling step; in fact, other ways involving SDS or proteinase K may affect adversely in PCR. The process may also be employed to amplify RNA, the process known as reverse transcriptase PCR (RT-PCR). This allows mRNA transcription products to be effectively analysed. It may also be used to differentiate latent viruses (detected by standard PCR) or active viruses, which replicate and thus produce transcription products and are, therefore, detectable by RT-PCR. In addition PCR may be extended to determine relative amount of transcription product.
Sensitivity of PCR:
The enormous sensitivity of PCR is also its one of the main drawbacks, since the very large degree of amplification makes the system vulnerable to contamination. Even a trace of foreign DNA, such as that contained in dust particles, may be amplified to significant levels and may mislead the results. Hence, cleanliness is a paramount when carrying out PCR and dedicated equipment and in some cases laboratories are used.
It is also possible that amplified products may also contaminate the PCR, although this may be overcome by ultraviolet irradiation to damage already amplified products so that they cannot be used as templates. Another way to overcome this kind of contamination is the use of uracil in the PCR and then to treat the products with enzyme N-glycocylase (UNG) which degrades any PCR product with incorporated uracil, rendering them useless to be used as templates.
The success of PCR process has given impetus to the development of other amplification based on either thermal or non-thermal (isothermal) cycling methods. The most popular alternative to the PCR is termed as “Ligase chain reaction or LCR”. This operates in a fashion similar to PCR but a thermo stable DNA ligase joins sets of primer together that are complimentary to target DNA.
ADVERTISEMENTS:
Following this, a similar exponential amplification reaction takes place, producing amounts of DNA that are similar to template DNA. Other alternative amplification methods are followed which all are based on one of the two strategies of either amplifying the target DNA/RNA molecule or by detecting the target and amplifying a signal molecule bound to it.
Target amplification method includes Ligase chain reaction (LCR) employing thermo stable DNA ligase used for mutation detection, etc. and other method under this category is nucleic acid sequence based amplification (NASBA) involving use of RNA, RNase H/reverse transcriptase, and T7 DNA polymerase used for viral detection, etc. (e.g., HIV).
Signal amplification method includes branched DNA amplification (β-DNA) involving isothermal micro-well format using hybridization or target /capture probe and signal amplification. It is used for mutation detection.
Factors Affecting the PCR:
1. Denaturing Temperature and Time:
The specific complementary association due to hydrogen bonding of single-stranded nucleic acids is referred to as “annealing”: two complementary sequences will form hydrogen bonds between their complementary bases (G to C, and A to T or U) and form a stable double-stranded, anti- parallel “hybrid” molecule.
ADVERTISEMENTS:
One may make nucleic acid (NA) single-stranded for the purpose of annealing, if it is not single-stranded already, like most RNA viruses, by heating it to a point above the “melting temperature” of the double- or partially-double-stranded form, and then flash-cooling it; this ensures the “denatured” or separated strands do not re-anneal. Additionally, if the NA is heated in buffers of ionic strength lower than 150 mM NaCl, the melting temperature is generally less than 100°C — which is why PCR works with denaturing temperatures of 91-97°C.
Taq polymerase is given as having a half-life of 30 min at 95°C, which is partly why one should not do more than about 30 amplification cycles. However, it is possible to reduce the denaturation temperature after about 10 rounds of amplification, as the mean length of target DNA is decreased. For templates of 300 bp or less, denaturation temperature may be reduced to as low as 88°C for 50% (G+C) templates, which means one may do as many as 40 cycles without much decrease in enzyme efficiency.
“Time at temperature” is the main reason for denaturation / loss of activity of Taq. Thus, if one reduces this, one will increase the number of cycles that are possible, whether the temperature is reduced or not. Normally the denaturation time is 1 min at 94°C. It is possible, for short template sequences, to reduce this to 30 sec or less. Increase in denaturation temperature and decrease in time may also work: recommend 96°C for 15 sec.
2. Annealing Temperature and Primer Design:
Primer length and sequence are of critical importance in designing the parameters of a successful amplification. The melting temperature of nucleic acid duplex increases both with its length, and with increasing (G+C) content. A simple formula for calculation of the Tm is
Tm = 4(G + C) + 2(A + T)°C.
Thus, the annealing temperature chosen for a PCR depends directly on length and composition of the primer(s). One should aim at using an annealing temperature (Ta) about 5°C below the lowest Tm of the pair of primers to be used. A more rigorous treatment of annealing temperature is given by Rychlik et al. (1990). They maintain that if the annealing temperature is increased by 1°C in every other cycle, specificity of amplification and yield of products <1kb in length are both increased.
ADVERTISEMENTS:
One consequence of having too low annealing temperature is that one or both primers will anneal to sequences other than the true target, as internal single-base mismatches or partial annealing may be tolerated. This is fine if one wishes to amplify similar or related targets. However, it can lead to “non-specific” amplification and consequent reduction in yield of the desired product, if the 3′-most base* is paired with a target.
A consequence of too high annealing temperature is that too little product will be made, as the likelihood of primer annealing is reduced; another and important consideration is that a pair of primers with very different annealing temperatures may never give appreciable yields of a unique product, and may also result in inadvertent “asymmetric” or single-strand amplification of the most efficiently primed product strand. Annealing does not take long, most primers will anneal efficiently in 30 sec or less, unless the annealing temperature (Ta) is too close to the Tm, or unless they are unusually long.
An illustration of the effect of annealing temperature on the specificity and on the yield of amplification of Human papillomavirus type 16 (HPV-16) is given below (Williamson and Rybicki, 1991: J Med Virol 33: 165-171).
3. Primer Length:
The optimum length of a primer depends upon its (A+T) content, and the Tm of its partner if one runs the risk of having problems as described above. Apart from the Tm, a prime consideration is that the primers should be complex enough so that the likelihood of annealing to sequences other than the chosen target is very low.
For example, there is a 1 /4 chance (4_1) of finding an A, G, C or T in any given DNA sequence; there is a 1/16 chance (4-2) of finding any dinucleotide sequence (e.g., AG); a 1/256 chance of finding a given 4-base sequence. Thus, a sixteen base sequence will statistically be present only once in every 416 bases (= 4 294 967 296 or 4 billion). This is about the size of the human or maize genome, and 1000 × greater than the genome size of E. coli.
ADVERTISEMENTS:
Thus, the association of a greater-than- 17-base oligonucleotide with its target sequence is an extremely sequence-specific process, far more so than the specificity of monoclonal antibodies in binding to specific antigenic determinants. Consequently, 17-mer or longer primers are routinely used for amplification from genomic DNA of animals and plants. Too long a primer length may mean that even high annealing temperatures are not enough to prevent mismatch pairing and non-specific priming.
4. Degenerate Primers:
For amplification of cognate sequences from different organisms, or for “evolutionary PCR”, one may increase the chances of getting product by designing “degenerate” primers. These would in fact be a set of primers which have a number of options at several positions in the sequence so as to allow annealing to and amplification of a variety of related sequences. For example, Compton (1990) described the use of 14-mer primer sets with 4 and 5 degeneracies as forward and reverse primers, respectively, for the amplification of glycoprotein B (gB) from related herpes viruses.
The reverse primer sequence was as follows:
TCGAATTCNCCYAAYTGNCCNT
where Y = T + C, and N = A + G + C + T, and the 8-base 5′-terminal extension comprises a EcoBI site (underlined) and flanking spacer to ensure that the restriction enzyme can cut the product (the New England Biolabs catalogue gives a good list of which enzymes require how long a flanking sequence in order to cut stub ends). Degeneracies obviously reduce the specificity of the primer(s), meaning mismatch opportunities are greater, and background noise increases; also, increased degeneracy means concentration of the individual primers decreases; thus, greater than 512-fold degeneracy should be avoided.
5. Elongation Temperature and Time:
This is normally 70-72°C, for 0.5-3 min. Taq actually has a specific activity at 37°C which is very close to that of the Klenow fragment of E. coli DNA polymerase I, which accounts for the apparent paradox which results when one tries to understand how primers which anneal at an optimum temperature can then be elongated at a considerably higher temperature, the answer is that elongation occurs from the moment of annealing, even if this is transient, which results in considerably greater stability.
At around 70°C the activity is optimal, and primer extension occurs at up to 100 bases/sec. About 1 min is sufficient for reliable amplification of 2 kb sequences. Longer products require longer times: 3 min is a good bet for 3 kb and longer products. Longer times may also be helpful in later cycles when product concentration exceeds enzyme concentration (>1 nM), and when dNTP and / or primer depletion may become limiting.
6. PCR Reaction Buffer:
Recommended buffers generally contain:
i. 10-50 mM Tris-HCl pH 8.3,
ii. Up to 50 mM KCl, 1,5mM or higher MgCl2,
iii. Primers 0.2-1 µM each primer,
iv. 50-200 µM each dNTP,
v. Gelatin or BSA to 100 µg/ml,
vi. And/or non-ionic detergents such as Tween-20 or Nonidet P-40 or Triton X-100 (0.05- 0.10% v/v)
Modern formulations may differ considerably; however, they are also generally proprietary. Following are some considerations which should be kept in mind during PCR.
(a) Higher than 50 mM KCl or NaCl inhibits Taq, but some is necessary to facilitate primer annealing.
(b) [Mg2+] affects primer annealing; Tm of template, product and primer-template associations; product specificity; enzyme activity and fidelity. Taq requires free Mg2+, so allowances should be made for dNTPs, primers and template, all of which chelate and sequester the cation; of these, dNTPs are the most concentrated, so [Mg2+] should be 0.5-2.5 mM greater than [dNTP]. A titration should be performed with varying [Mg2+] with all new template-primer combinations, as these can differ markedly in their requirements, even under the same conditions of concentrations and cycling times/temperatures.
(c) Some enzymes do not need added protein, others are dependent on it. Some enzymes work markedly better in the presence of detergent, probably because it prevents the natural tendency of the enzyme to aggregate.
(d) Primer concentrations should not go above 1 µM unless there is a high degree of degeneracy; 0.2 (J.M is sufficient for homologous primers.
(e) Nucleotide concentration need not be above 50 µM each; long products may require more, however.
7. Cycle Number:
The number of amplification cycles necessary to produce a band visible on a gel depends largely on the starting concentration of the target DNA. Innis and Gelfand (1990) recommend from 40-45 cycles to amplify 50 target molecules, and 25-30 to amplify 3×105 molecules to the same concentration. This non-proportionality is due to a so-called plateau effect, which is the attenuation in the exponential rate of product accumulation in late stages of a PCR, when product reaches 0.3-1.0 nM.
This may be caused by degradation of reactants (dNTPs, enzyme); reactant depletion (primers, dNTPs-former a problem with short products, latter for long products); end-product inhibition (pyrophosphate formation); competition for reactants by non-specific products; competition for primer binding by re-annealing of concentrated (10 nM) product.
If desired product is not made in 30 cycles, take a small sample (1 ul) of the amplified mix and re-amplify 20-30x in a new reaction mix rather than extending the run to more cycles. In some cases where template concentration is limiting, this can give good product where extension of cycling to 40x or more does not.
A variant of this is nested primer PCR. PCR amplification is performed with one set of primers, and then some product is taken, with or without removal of reagents, for re-amplification with an internally-situated, “nested” set of primers. This process adds another level of specificity, meaning that all products non-specifically amplified in the first round will not be amplified in the second. This is illustrated in the left side:
8. Helix De-stabilisers / Additives:
With nucleic acids of high (G+C) content, it may be necessary to use harsher denaturation conditions. For example, one may incorporate up to 10% (w or v/v):
i. Dimethyl sulphoxide (DMSO),
ii. Dimethyl form amide (DMF),
iii. Urea or
iv. Form amide
In the reaction mixture these additives are presumed to lower the Tm of the target nucleic acids, although DMSO at 10% and higher is known to decrease the activity of Taq by up to 50%. Additives may also be necessary in the amplification of long target sequences. DMSO often helps in amplifying products of >1kb.
Form amide can apparently dramatically improve the specificity of PCR, while glycerol improves the amplification of high (G+C) templates. Polyethylene glycol (PEG) may be a useful additive when DNA template concentration is very low. It promotes macromolecular association by solvent exclusion, meaning the pol (DNA polymerase enzyme) can find the DNA.
Applications for PCR:
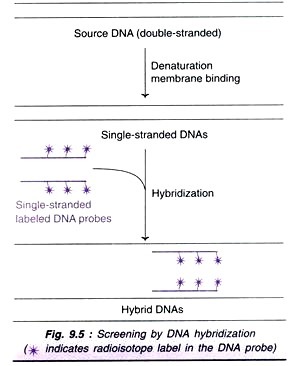
1. The labelling of gene probes is one such area which has traditionally been undertaken by techniques such as nick translation. The nature of PCR makes it an ideal method for gene probe production and labelling.
2. Important modification of PCR known as quantitative PCR by which the initial concentration of template DNA can be estimated and is very useful for the measurement of, for example, a virus or an mRNA for protein expressed in abnormal amount in a disease process. Early quantitative PCR methods involved the comparison of a standard or control DNA template amplified with separate primers at the same time as the specific target DNA.
These kind of quantification rely on the reaction being exponential and so any factor affecting this may also affect the result. Other method involves the incorporation of radiolabel through primer and their detection in subsequent purification of PCR products.
3. One of the most general applications of development of PCR is direct PCR sequencing. This traditionally involves the cloning of sequences into vectors developed for chain termination sequencing. However, rapid accumulation of PCR products allows nucleotide sequence information to be obtained very quickly.
Variations on the Basic PCR Technique:
1. Allele-specific PCR:
AS-PCR is used to determine the genotype of single-nucleotide polymorphisms (SNPs) (single base differences in DNA) by using primers whose ends overlap the SNP and differ by that single base. PCR amplification is less efficient in the presence of a mismatch, so the differences in amplification resulting from different primers can be used to quickly determine which primer matches the sample genotype.
2. Assembly PCR:
Assembly PCR is the completely artificial synthesis of long gene products by performing PCR on a pool of long oligonucleotides with short overlapping segments. The oligonucleotides alternate between sense and antisense directions and the overlapping segments serve to order the PCR fragments so that they selectively produce their final product.
3. Asymmetric PCR:
Asymmetric PCR is used to preferentially amplify one strand of the original DNA more than the other. It finds use in some types of sequencing and hybridization probing where having only one of the two complementary stands is required. PCR is carried out as usual, but with a great excess of the primers for the chosen strand.
Due to the slow (arithmetic) amplification later in the reaction after the limiting primer has been used up, extra cycles of PCR are required. A recent modification on this process, known as Linear-After-The-Exponential-PCR (LATE-PCR), uses a limiting primer with a higher melting temperature (TJ than the excess primer to maintain reaction efficiency as the limiting primer concentration decreases mid-reaction.
4. Colony PCR:
Bacterial clones (E. coli) can be rapidly screened for correct DNA vector constructs. Selected bacterial colonies are picked with a sterile toothpick from an agarose plate and dabbed into the master mix or sterile water. Primers (and the master mix) are added, and the PCR is started with an extended time at 95°C when standard polymerase is used or with a shortened denaturation step at 100°C and special chimeric DNA polymerase.
5. Helicase-dependent amplification:
Similar to traditional PCR, but maintains a constant temperature rather than cycling through denaturation and annealing/extension cycles. Helicase, an enzyme that unwinds DNA, is used in place of thermal denaturation.
6. Hot-start PCR:
Hot-start PCR is a technique that reduces non-specific amplification during the initial set up stages of the PCR. The technique may be performed manually by simply heating the reaction components briefly at the melting temperature (e.g., 95°C) before adding the polymerase.
Specialized enzyme systems have been developed that inhibit the polymerase’s activity at ambient temperature, either by the binding of an antibody or by the presence of covalently bound inhibitors that only dissociate after a high-temperature activation step. Hot-start/cold-finish PCR is achieved with new hybrid polymerases that are inactive at ambient temperature and are instantly activated at elongation temperature.
7. Inter-sequence specific (ISSR) PCR:
A PCR method for DNA fingerprinting that amplifies regions between some simple sequence repeats to produce a unique fingerprint of amplified fragment lengths.
8. Inverse PCR:
Inverse PCR is a method used to allow PCR when only one internal sequence is known. This is especially useful in identifying flanking sequences to various genomic inserts. This involves a series of DNA digestions and self-ligation, resulting in known sequences at either end of the unknown sequence.
9. Ligation-mediated PCR.
10. Methylation Specific PCR:
Methylation Specific PCR (MSP) is used to detect methylation of CpG islands in genomic DNA. DNA is first treated with sodium bi-sulphite, which converts un-methylated cytosine bases to uracil, which is recognized by PCR primers as thymine. Two PCR reactions are then carried out on the modified DNA, using primer sets identical except at any CpG islands within the primer sequences.
At these points, one primer set recognizes DNA with cytosine’s to amplify methylated DNA, and one set recognizes DNA with uracil or thymine to amplify un-methylated DNA. MSP using qPCR can also be performed to obtain quantitative rather than qualitative information about methylation.
11. Multiplex Ligation-dependent Probe Amplification (MLPA):
Permits multiple targets to be amplified with only a single primer pair, thus avoiding the resolution limitations of multiplex PCR (see below).
12. Multiplex-PCR:
ADVERTISEMENTS:
The use of multiple, unique primer sets within a single PCR reaction to produce amplicons of varying sizes specific to different DNA sequences. By targeting multiple genes at once, additional information may be gained from a single test run that otherwise would require several times the reagents and more time to perform.
Annealing temperatures for each of the primer sets must be optimized to work correctly within a single reaction, and amplicon sizes, i.e., their base pair length, should be different enough to form distinct bands when visualized by gel electrophoresis.
13. Nested PCR:
Nested PCR increases the specificity of DNA amplification, by reducing background due to non-specific amplification of DNA. Two sets of primers are being used in two successive PCR reactions. In the first reaction, one pair of primers is used to generate DNA products, which besides the intended target, may still consist of non- specifically amplified DNA fragments.
The product(s) (sometimes after gel purification after electrophoresis of the PCR product) are then used in a second PCR reaction with a set of primers whose binding sites are completely or partially different from the primer pair used in the first reaction, but are completely within the DNA target fragment. Nested PCR is often more successful in specifically amplifying long DNA fragments than conventional PCR, but it requires more detailed knowledge of the target sequences.
14. Quantitative PCR:
Q-PCR (Quantitative PCR) is used to measure the quantity of a PCR product (preferably real-time). It is the method of choice to quantitatively measure starting amounts of DNA, cDNA or RNA. Q-PCR is commonly used to determine whether a DNA sequence is present in a sample and the number of its copies in the sample.
The method with currently the highest level of accuracy is Quantitative real-time PCR. It is often confusingly known as RT-PCR (Real Time PCR) or RQ-PCR. QRT-PCR or RTQ-PCR is more appropriate contractions. RT-PCR commonly refers to reverse transcription PCR (see below), which is often used in conjunction with Q- PCR. QRT-PCR methods use fluorescent dyes, such as Sybr Green, or fluorophore- containing DNA probes, such as TaqMan, to measure the amount of amplified product in real time.
15. RT-PCR:
RT-PCR (Reverse Transcription PCR) is a method used to amplify, isolate or identify a known sequence from a cellular or tissue RNA. The PCR reaction is preceded by a reaction using reverse transcriptase to convert RNA to cDNA.
RT-PCR is widely used in expression profiling, to determine the expression of a gene or to identify the sequence of an RNA transcript, including transcription start and termination sites and, if the genomic DNA sequence of a gene is known, to map the location of exons and introns in the gene. The 5′ end of a gene (corresponding to the transcription start site) is typically identified by a RT-PCR method, named RACE-PCR, short for Rapid Amplification of cDNA Ends.
16. TAIL-PCR:
Thermal asymmetric interlaced PCR is used to isolate unknown sequence flanking a known sequence. Within the known sequence TAIL-PCR uses a nested pair of primers with differing annealing temperatures; a degenerate primer is used to amplify in the other direction from the unknown sequence.
17. Touchdown PCR:
Touchdown PCR is a variant of PCR that aims to reduce nonspecific background by gradually lowering the annealing temperature as PCR cycling progresses. The annealing temperature at the initial cycles is usually a few degrees above the Tm of the primers used, while at the later cycles, it is a few degrees below the primer Tm. The higher temperatures give greater specificity for primer binding, and the lower temperatures permit more efficient amplification from the specific products formed during the initial cycle.