ADVERTISEMENTS:
Read this article to learn about the basis, isolation and limitations of somaclonal variations and also factors affecting the production of somaclonal variants.
The genetic variations found in the in vitro cultured cells are collectively referred to as somaclonal variations.
The plants derived from such cells are referred to somaclones. Some authors use the terms calliclones and proto-clones to represent cultures obtained from callus and protoplasts respectively.
ADVERTISEMENTS:
The growth of plant cells in vitro is an asexual process involving only mitotic division of cells. Thus, culturing of cells is the method to clone a particular genotype. It is therefore expected that plants arising from a given tissue culture should be the exact copies of the parental plant.
The occurrence of phenotypic variants among the regenerated plants (from tissue cultures) has been known for several years. These variations were earlier dismissed as tissue culture artefacts. The term somaclonal variations was first used by Larkin and Scowcraft (1981) for variations arising due to culture of cells, i.e., variability generated by a tissue culture. This term is now universally accepted.
As described elsewhere the explant used in tissue culture may come from any part of the plant organs or cells. These include leaves, roots, protoplasts, microspores and embryos. Somaclonal variations are reported in all types of plant tissue cultures.
In recent years, the term gametoclonal variations is used for the variations observed in the regenerated plants from gametic cells (e.g., anther cultures). For the plants obtained from protoplast cultures, proto-clonal variations is used.
ADVERTISEMENTS:
Basis of Somaclonal Variations:
Somaclonal variations occur as a result of genetic heterogeneity (change in chromosome number and/or structure) in plant tissue cultures.
This may be due to:
i. Expression of chromosomal mosaicism or genetic disorders.
ii. Spontaneous mutations due to culture conditions.
The genetic changes associated with somaclonal variations include polyploidy, aneuploidy, chromosomal breakage, deletion, translocation, and gene amplifications, besides several mutations. In fact, the presence of several chromosomal aberrations—reciprocal translocation, deletions, inversions, chromosomal breakage, multi-centric, acentric fragments have been found among the somaclones of barley, garlic and oat.
The occurrence of mutations in cultures is relatively low. Mutations may be due varied nutrients, culture conditions and mutagenic effects of metabolic products that accumulate in the medium. Somaclonal variations due to transposable elements, mitotic crossing over and changes in the cytoplasmic genome have also been reported.
Nomenclature of somaclones:
The somaclones that are regenerated from tissue cultures directly are regarded as R0 or R plants. The self-fertilized progeny of R0 plants represent R1 plants. R2, R3, R4 etc. plants are the subsequent generations. Some workers use other nomenclature — somaclones (SC1 = R0), SC2, SC3, SC4 etc. for subsequent generations.
Isolation of Somaclonal Variants:
There are two procedures commonly used for obtaining the crop plants with somaclonal variations:
ADVERTISEMENTS:
1. Without in vitro selection
2. Within vitro selection.
1. Without in Vitro Selection:
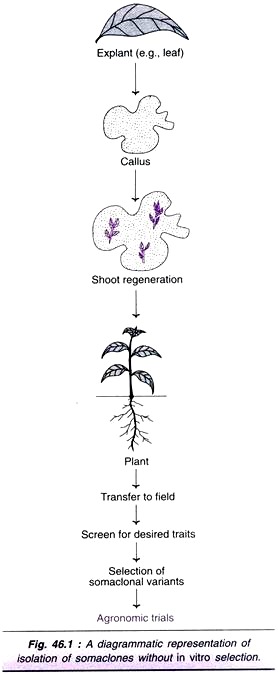
An explant (leaf, stem, root etc.) is cultured on a suitable medium, supplemented with growth regulators. The unorganized callus and cells do not contain any selective agent (toxic or inhibitory substance). These cultures are normally sub-cultured, and transferred to shoot induction medium for regeneration of plants. The so produced plants are grown in pots, transferred to field, and analyzed for somaclonal variants (Fig. 46.1).
Somaclonal variants of several crops have been successfully obtained by this approach e.g., sugarcane, potato, tomato, cereals etc. A selected list of disease resistant crop plants obtained from somaclonal variations without in vitro selection along with the pathogenic organisms is given in Table 46.1.
Limitations of without in vitro selection approach:
There is no directed and specific approach for the isolation of somaclones without in vitro selection. Consequently, the appearance of a desired trait is purely by chance. Further, this procedure is time consuming and requires screening of many plants.
2. With in Vitro Selection:
ADVERTISEMENTS:
Isolation of somaclones with in vitro selection method basically involves handling of plant cells in cultures (protoplast, callus) like microorganisms and selection of biochemical mutants. The cell lines are screened from plant cultures for their ability to survive in the presence of a toxic/inhibitory substance in the medium or under conditions of environmental stress.
A diagrammatic representation of in vitro protocol for the isolation of disease resistance plants with in vitro selection approach is given in Fig. 46.2.
The differentiated callus, obtained from an explant is exposed in the medium to inhibitors like toxins, antibiotics, amino acid analogs. Selection cycles are carried out to isolate the tolerant callus cultures and these calli are regenerated into plants. The plants so obtained are in vitro screened against the toxin (or pathogen or any other inhibitor).
ADVERTISEMENTS:
The plants resistant to the toxin are selected and grown further by vegetative propagation or self-pollination. The subsequent generations are analysed for disease resistant plants against the specific pathogenic organism.
A selected list of disease resistant crop plants obtained from somaclonal variations by in vitro selection along with pathogenic organisms and selection agents is given in Table 46.2.
Besides the disease resistant plants, plants with herbicide resistance and antibiotic resistance have also been developed with in vitro selection approach.
Environmental stress tolerance:
High salt concentration in the soil is the major constraint limiting the crop development and yield. Many plants with salt tolerance (salinity) have been developed e.g. rice, tobacco. Several attempts are being made by plant biotechnologists to develop cold tolerant crops, although the success rate has been very limited.
ADVERTISEMENTS:
Advantages of with in vitro selection approach:
The major advantage of with in vitro selection method is the specific selection of the desired trait rather than a general variation found at the plant level. This procedure is less time consuming when compared to without in vitro selection approach.
Factors Affecting Production of Somaclonal Variants:
Some of the important factors that influence development of somaclonal variants by without in vitro selection and within vitro selection are briefly described.
Genotype and explant source:
The nature of genotype of the plants influences the frequency of regeneration and frequency of production of somaclones. Explants can be taken from any part of plant — leaves, roots, internodes, ovaries etc. The source of explant is very critical for somaclonal variations. For instance, potato plants regenerated from callus of rachis and petiole are much higher (~50%) compared to those regenerated from callus of leaves (~12%).
Duration of cell culture:
ADVERTISEMENTS:
In general, for many plant cultures, somaclonal variations are higher with increased duration of cultures. For example, it was reported that genetic variability increased in tobacco protoplasts from 1.5 to 6% by doubling the duration of cultures.
Growth hormone effects:
The plant growth regulators in the medium will influence the karyotypic alterations in qultured cells, and therefore development of somaclones. Growth hormones such as 2, 4-dichlorophenoxy acetic acid (2, 4-D) and naphthalene acetic acid (NAA) are frequently used to achieve chromosomal variability.
Besides the factors discussed above, selection of somaclones with in vitro selection are dependent on the following parameters:
i. Selection propagule (cells, protoplasts, calli).
ii. Selection agent (toxin, herbicide, amino acid analogue).
iii. Technique used for selection.
iv. Stability of resistant substance.
v. In vivo testing procedure.
vi. Ability for regeneration of plants.
Limitations of Somaclonal Variations:
Despite several applications of somaclonal variations, there are certain limitations/ disadvantages also:
i. Most of the somaclonal variations may not be useful.
ii. The variations occur in an unpredictable and uncontrolled manner.
iii. Many a times the genetic traits obtained by somaclonal variations are not stable and heritable.
iv. Somaclonal variations are cultivar-dependent which is frequently a time consuming process.
v. Somaclones can be produced in only those species which regenerate to complete plants.
vi. Many cell lines (calli) may not exhibit regeneration potential.
Gametoclonal Variations:
The variations observed while culturing the gametic cells are regarded as gametoclonal variations. This is in contrast to the somaclonal variations detected in the cultures of somatic tissues.
The term gametoclones (in place of somaclones) is used for the products of gametoclonal variations.
As the somatic cells divide by mitosis, the genetic material is equally distributed to the daughter cells. In contrast, the gametes, being the products of meiosis, possess only half of the parent cell genetic material.
The gametoclonal variations differ from somaclonal variations by three distinct features:
1. Mutants obtained from gametoclonal variations give rise to haploid plants since a single set of chromosomes are present.
2. Meiotic crossing over is the recombination process observed in gametoclonal variations.
3. The gametoclones can be stabilized by doubling the chromosome number.
Production of Gametoclones:
Gametoclones can be developed by culturing male or female gametic cells. The cultures of anthers or isolated microspores are widely used. A selected list of plants regenerated from gametoclonal variations is given in Table 46.4. Improvements have been made in several plant species through development of gametoclones e.g., rice, wheat, and tobacco.
Source of Gametoclonal Variations:
There are three sources of genetic variations in the gametoclones:
1. Cell culture technique may induce genetic variations.
2. Variations may be induced while doubling the haploid chromosomes.
3. Genetic variations may occur due to heterozygosity of the diploids.