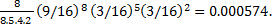ADVERTISEMENTS:
Are you looking for an essay on ‘Genetic Code’? Find paragraphs, long and short essays on ‘Genetic Code’ especially written for college and medical students.
Essay on Genetic Code
Essay Contents:
- Essay on the Concept of Genetic Code
- Essay on the Methods Used for Cracking the Genetic Code
- Essay on the Characteristics of Triplet Codon
- Essay on the Wobble Hypothesis on Genetic Code
ADVERTISEMENTS:
Essay # 1. Concept of Genetic Code:
ADVERTISEMENTS:
Amino acid sequences of the protein are governed by the nucleotide sequences of the mRNA. This specific relationship between the nucleotide sequence and amino acid sequence is known as genetic code, which was deciphered by Marshall Nirenberg and his colleagues in early 1960s.
DNA transfers information to mRNA in the form of a code defined by a sequence of nucleotides bases. During protein synthesis, ribosomes move along the mRNA molecule and “read” its sequence three nucleotides at a time (codon) from the 5′ end to the 3′ end. Each amino acid is specified by the mRNA’s codon and then pairs with a sequence of three complementary nucleotides carried by a particular tRNA (anticodon). The genetic code consists of 64 triplets of nucleotides (Fig. 8.2). These triplets are called codons.
Since RNA is constructed from four types of nucleotides, there are 64 possible triplet sequences or codons (4x4x4=64). With three exceptions, each codon encodes for one of the 20 amino acids used in the synthesis of proteins that produces some redundancy in the code- most of the amino acids being encoded by more than one codon. Three of these possible codons specify the termination of the polypeptide chain, called “stop codons”, that leaves 61 codons to specify only 20 different amino acids. Therefore, most of the amino acids are represented by more than one codon. The genetic code is said to be degenerate.
ADVERTISEMENTS:
The genetic code can be expressed as either RNA codons or DNA codons. RNA codons occur in messenger RNA (mRNA) and are the codons that are actually “read” during the synthesis of polypeptides (the process called translation). But each mRNA molecule acquires its sequence of nucleotides by transcription from the corresponding gene.
Essay # 2. Methods Used for Cracking the Genetic Code:
DNA is a genetic material, which carries genetic information from cell to cell and from generation to generation. The genetic information may be written in any one of the three moieties of DNA, phosphoric acid, deoxyribose sugar, and nitrogen bases. But the poly-sugar- phosphate backbone is always the same, and it is therefore, unlikely that these moieties of DNA molecule carry the genetic information. The nitrogen bases, however vary from one segment of DNA to another, so the information might well depend on their sequences. The sequences of nitrogen bases of a given segment of DNA molecule, actually has been found to be identical to linear sequence of amino acids in a protein molecule.
The proof of such colinearity between DNA nitrogen base sequence and amino acid sequence in protein molecules has first obtained from an analysis of mutants of head protein of bacteriophage T4, and the A-protein tryptophan synthetase of Escherichia coli. The colinearity of protein molecules and DNA polynucleotides has given the clue that the specific arrangement of four nitrogen bases (e.g., A, T, C and G) in DNA polynucleotide chains somehow, determines the sequence of amino acids in protein-molecules. Therefore, these four DNA bases can be considered as four alphabets of DNA molecule.
All the genetic information, therefore, should be written by these four alphabets of DNA. The genetic informations were supposed to be existed in DNA molecule in the form of certain special language of code words which might utilized the four nitrogen bases of DNA for its symbols.
The different methods used for cryptoanalysis or cracking the genetic code are:
a. In Vitro Codon Assignments:
Nirenberg in 1961, first of all, provided the means for a much more rapid decipherment of the genetic code. He made in vitro studies on the incorporation of radioactive amino acids in cell free protein synthetic systems containing artificial (synthetic) ribopolynucleotides or mRNA. A cell free system for protein synthesis includes ribosomes, enzymes, tRNA, etc. To produce such a cell free protein synthetic system the protein synthesizing components of broken cell (usually E. coli) were separated from the remainder by gentle centrifugation. During these manipulations natural mRNA is subjected to mechanical and enzymatic break down. Artificial messenger RNAs were then introduced.
ADVERTISEMENTS:
The artificial mRNA was prepared for each nitrogen base with the aid of a polynucleotide phosphorylase enzyme obtained from Azobacter vinelandii or Micrococcus lysodeikticus. The enzyme differed from RNA polymerase enzyme in not requiring the presence of DNA as a primer or template for the synthesis of RNA. Thus, Nirenberg prepared a polyu- ridylic acid (UUUU …), poly cytidylic acid (CCCCC …), polyadenylic acid (AAAAA … ) and guanylic acid (GGGGG …) from uracil, cytosine, adenine and guanine respectively. When he introduced these artificial homopolymeric polyribonucleotides or mRNA (e.g., poly D, poly C, poly A and poly G acids) in different cell free protein synthetic systems of E. coli containing radioactive amino acids, he found that each artificial mRNA stimulated the synthesis of polypeptides of single kind of amino acids.
For example, poly U acid stimulates the in vitro synthesis of monotonus polypeptide-namely polyphenylalanine whose amino acid residues were phenylalanine. Therefore, he concluded that a sequence of UUU coded for phenylalanine. Likewise, because polycytidylic acid was found to direct synthesis of polyproline hence, the sequence of CCC codes designates proline and because, polyadenylic acid directs synthesis of poly lysine, so the sequence of AAA designates lysine. Later on, the sequence of GGG was found to specify glycine.
In this way four of 64 codons were easily accounted for. In order to gain some insight into the meaning of the remaining 60 codons, containing more than one kind of nucleotide Nirenberg continued these experiments by using artificially synthesized random ribopolynucleotides (mixed copolymers) containing two, three or four different nucleotide constituents. By using artificial mRNAs, viz., poly-UA, poly UC, and poly UG, he cracked the codes for arginine, alanine, serine, proline, tyrosine, isoleucine, valine, leucine, cysteine, tryptophan, glycine, methionine and glutamic acid.
Further, when mixed copolymers are made, one can calculate all the possible triplets they contain assuming that the bases are incorporated randomly into the molecule; for example, when poly AC is synthesized from a mixture containing equal proportions of A and C, six triplets should occur with equal frequency- AAA, ACA, ACC, CCC, CCA and CAA. Using poly AC it was found that six amino acids were incorporated into polypeptides- aspargine, glutamine, histidine, lysine, threonine and proline.
ADVERTISEMENTS:
The next step was to vary the ratio of A and C, and it was found that when the copolymer contained more A than C the ratio of aspargine to histidine in the polypeptide increased, By such experiments the composition of the bases in certain triplets could often be deduced.
After learning of Nirenberg’s first success with poly U, Ochoa and his collaborators carried out entirely analogous sets of decoding experiments in which they used artificial random ribopolynucleotides. Through the parallel efforts of these two research groups the empirical formula of many amino acid codes or codons became known within a year or so. But this- work could not establish the actual sequence of the nucleotides within any codon, except for, UUU, CCC, AAA and GGG.
In order to ascertain that which structural isomer of a codon of a given empirical formula designates which amino acid, artificial mRNAs of known sequences were employed. H. G. Khorana first of all succeeded in synthesizing polyribonucleotides or mRNA of known sequences and be synthesized an alternating copolymer such as-UGU-GUG-UGU-.
Use of this particular alternating polynucleotide as a mRNA in the in vitro protein-synthesizing system gave rise to the formation of the alternating polypeptide (e.g. valine-cysteine valine-cysteine) evidently directed by the codon sequence-UGU-GUG-UGU-GUG. This result established that the UUG (Iucine) codon is not UGU, and that neither the UUG (tryptophan) nor the UGG (glycine) codon is GUG, but, UUG (valine) is UGU, and GUG is a cysteine synonym, or that UUG (cysteine) is UGU nor GUG is a valine synonym. Thus, Khorana determined the exact sequence of nucleotides in various genetic codons which discovered either by Nirenberg or by himself.
ADVERTISEMENTS:
In all of these in vitro experiments magnesium concentrations were kept artificially high in order to facilitate the translation of artificial messengers. High magnesium concentrations allowed initiation without an initiation codon, and an AUG codon is, of course, absent from most of the artificial polyribonucleotides that might be synthesized in vitro.
In 1964, while Khorana was just completing the difficult synthesis of the first artificial ribopolynucleotides of defined sequence, Nirenberg made a second discovery that made possible a rapid culmination of the code-breaking efforts. He found that addition of simple trinucleotides of known base sequences to ribosomes would cause these ribosomes to bind that, and only that, aminoacyl-tRNA which carries the anticodon complementary to the trinucleotide added to the reaction mixture. For example, the trinucleotide GCC was found to be active in promoting the binding of alanyl-tRNAala but not of any other aminoacyl-tRNA and it was therefore concluded that GCC is a codon for alanine. These studies enabled the preparation of the complete genetic code dictionary.
b. In Vivo Codon Assignment:
The cell free protein synthetic systems, though have proved of great significance in decipherment of the genetic code, but, they could not tell us whether the genetic code so decipherd is used in the living systems of all organisms also.
ADVERTISEMENTS:
Three kinds of techniques are used by different molecular biologists to determine whether the same code is also used in vivo:
(a) Amino acid replacement studies (e.g., tryptophan synthetase synthesis in E. coli and haemoglobin synthesis in man),
(b) Frame shift mutations, on lysozyme enzyme of T4, bacteriophages, and
(c) Comparison of a DNA or mRNA polynucleotide cryptogram with its corresponding polypeptide clear text e.g., comparison of amino acid sequence of the R17 bacteriophage coat protein with the nucleotide sequence of the R17 mRNA in the region of the molecule that dictates coat- protein synthesis. Thus, in vitro and in vivo studies gave the way to formulate a code table for twenty amino acids. In genetic dictionary, each codon is written as it would appear in RNA reading in a 5’—>3′ direction; the corresponding codons in DNA will of course, be both complementary to them and written in the reverse order on a 5’—>3′ strand. Similarly, the bases in the corresponding tRNA anticodons will be both complementary and anti-polar to the mRNA codons.
Essay # 3. Characteristics of Triplet Codon:
The genetic dictionary of mRNA codons reveals following important features of triplet codons:
ADVERTISEMENTS:
a. Codon is Triplet:
Twenty different amino acids are incorporated during translation. Thus, at least 20 different codons must be formed using the four symbols (bases) available in the mRNA. Two bases per codon would yield only 42 or 16 possible codons which are clearly not enough. Three bases per codon yield 43 or 64 possible codons apparently excess. It is now known that each codon consists of a sequence of three nucleotides, i.e., it is a triplet code. The deciphering of the genetic code, depended heavily on the chemical synthesis of nucleotide polymer, particularly triplets in repeated sequence.
b. Degeneracy:
The code contains many synonyms, in that almost all amino acids are represented by more than one codon. The occurrence of more than one codon per amino acid is called degeneracy. All amino acids except methionine and tryptophan have more than one codon, so that all the possible triplets have a meaning, despite there being 64 triplets and only 20 amino acids. Leucine, serine and arginine have six different codons. Isoleucine has three codons. The degeneracy in the genetic code is not at random: instead, it is highly ordered. Usually multiple codons specifying an amino acid differ by only one base, the third or 3 base of codon.
For example, the three amino acids-arginine, serine and leucine-each have six synonymous codons. However, for many of the synonym codons specifying the same amino acid the first two bases of the triplet are constant, whereas the third can vary; for example, all codons starting with CC specify proline (CCU, CCC, CCA and CCG) and all codons starting with AC specify threonine. This flexibility in the third nucleotide of a codon may well help to minimize the consequences of errors.
The degeneracy can be either partial or complete. A complete degeneracy is a condition in which the third base position is of less significance. Any base present in this position will lead to same amino acid, but if the third position is occupied by one type of purine/ pyrimidine, then they code for one type of amino acid and if the position is occupied by another type of purine/ pyrimidines, without any change in the first two positions, they code for another amino acid.
ADVERTISEMENTS:
Such a condition is called as partial degeneracy, e.g. CAU or CAC code for His whereas CAA and CAG code for Gly. Because of the degeneracy of the genetic code there must either be several different tRNAs that recognize the different codons specifying a given amino acid or the anticodon of a given tRNA must be able to base-pair with several different codons. Actually both of these occur. Several tRNAs exist for certain amino acids and some tRNAs recognize more than one codon. The hydrogen bonding between the bases in the anticodon of tRNA and the codon of mRNA appears to follow strict base pairing rules only for the first two bases of the codon and is apparently less stringent, allowing what Crick has called wobble at this site.
c. Non-Overlapping:
The code is non-overlapping, meaning that no single base can take part in the formation of more than one codon. This means that successive triplets are read in order. Each nucleotide is part of only one triplet codon.
d. Ambiguity:
The genetic code is ambiguous, that is, same codon may specify more than one amino acid. For example, UUU codon usually code for phenylalanine but in the presence of streptomycin, may also code for isoleucine, leucine or serine.
e. Commaless:
The genetic code is commaless, which means that no codon is reserved for punctuations. There are no commas or some specific nucleotide sequences to separate the codons, i.e., CCCAAAUUUGGG have four code words and upon translation we have a tetrapeptide chain of pro-lys-phen-gly. So all the letters are used to code for one or other amino acid.
f. Starting Codons:
ADVERTISEMENTS:
AUG codon is called starting or chain initiation codon, because, it initiates, the synthesis of polypeptide chain.
g. Stop Codons/Non-Sense Codons:
There are 3 codons (UAA, UAG, and UGA) that are stop codons. When any one of these codons is reached during transcription, transcription stops. The UAA (also called ochre), UAG (also called amber because an investigator who studied the properties of this codon belonged to the Bernstein family, and Bernstein means “amber” in German) and UGA codons do not specify any amino acid, and so, are called nonsense codons. They are also called termination codons, because, these codons are used by the cell to signal the natural end of translation of a particular polypeptide. However, their inclusion in any mRNA results in the abrupt termination of the message at the point of their location even though the polypeptide chain has not been-completed.
h. Universality:
The genetic code has been found to be universal, because, same code applies in all kinds of living systems. It was postulated that the genetic code must be frozen and unable to evolve because a change in a codon meaning would cause almost every protein in the cell to be altered. The genetic codes are indeed universal and can be demonstrated quite directly by presenting an E. coli in in vitro protein synthesizing system with, for example, purified mRNA from polio virus (which is normally translated by human cells) and observing the synthesis of virus protein.
Experiments with cell free system with mammalian extract yielded same results as that of E. coli. The major exception to the universality of the code occurs in mitochondria of humans, yeast and several other species where UGA is a tryptophan codon. UGA is a termination codon in the non-mitochondrial system.
Also, in yeast mitochondria, CUA specifies threonine instead of the usual leucine and in mammalian mitochondria; AUA specifies methionine instead of isoleucine. Still more surprising is the discovery that the meaning of a codon can vary from gene to gene in the same organism. In human nuclear genes UGA is frequently used as a termination codon in accordance with its standard meaning. However, in at least two human genes, for the enzymes glutathione peroxidase and iodothyronine-5-deiodinase, UGA specifies the unusual amino acid called selenocysteine (a cysteine in which the sulphur atom is replaced by selenium).
It appears that the mRNA transcribed from these genes have special stem loop structures in their trailer regions, and these stem loops play some role during translation, ensuring that the UGA triplet is recognized as a selenocysteine codon rather than a termination signal.
Essay # 4. Wobble Hypothesis on Genetic Code:
The triplet code is a degenerate one with many more codons than the number of amino acid types coded. An explanation for this degeneracy is provided by the ‘wobble hypothesis’ proposed by Crick (1966). Since there are 61 codons specifying amino acids, the cell should contain 61 different tRNA molecules, each with a different anticodon. Actually, however, the number of tRNA molecule types discovered is much less than 61. This implies that the anticodons of some tRNAs read more than one codon on mRNA. According to the wobble hypothesis only the first two positions of a triplet codon on mRNA have a precise pairing with the bases of the tRNA anticodon.
The pairing of the third position bases of the codon may be ambiguous, and varies according to the nucleotide present in this position. Thus a single tRNA type is able to recognize two or more codons differing only in the third base. The anticodon UCG of serine tRNA recognizes two codons, AGC and AGU. The bonding between UCG and AGC follows the usual Watson-Crick pairing pattern. In UCG-AGU pairing, however, hydrogen bonding takes place between G and U. This is a departure from the usual Watson-Crick pairing mechanism where G pairs with C and A with U. Such interaction between the third bases is referred to as ‘wobble pairing’.
The degeneracy of the code is not random. Mostly, the different codons for a particular amino acid have the same first two letters (leucine, serine and arginine are exceptions). Thus the first two letters of all the four codons for valine are GU and for alanine GC.
Codon Bias:
All but two of the amino acids (Met and Trp) can be encoded by from 2 to 6 different codons. However, the genome of most organisms reveals that certain codons are preferred over others. In humans, for example, alanine is encoded by GCC four times as often as by GCG. Why this is uncertain? It probably reflects a greater translation efficiency by the translation apparatus (e.g., ribosomes) for certain codons over their synonyms.
Codon bias even extends to pairs of codons: wherever a human protein contains the amino acids Ala-Glu, the gene encoding those amino acids is seven times as likely to use the codons GCAGAG rather than the synonymous GCCGAA. Codon bias is exploited by the biotechnology industry to improve the yield of the desired product. The ability to manipulate codon bias may also usher in an era of safer vaccines.