ADVERTISEMENTS:
Here is a term paper on ‘Chromosomes’ for class 9, 10, 11 and 12. Find paragraphs, long and short term papers on ‘Chromosomes’ especially written for school and college students.
Term Paper on Chromosomes
Term Paper Contents:
- Term Paper on the Introduction to Chromosomes
- Term Paper on the Prokaryotic Chromosomes
- Term Paper on the Eukaryotic Chromosomes
- Term Paper on the DNA Molecules in Mitochondria
- Term Paper on the DNA Molecules in Chloroplasts
ADVERTISEMENTS:
1. Term Paper on the Introduction to Chromosomes:
ADVERTISEMENTS:
Chromosomes as the physical basis for the genetic properties of cells and for inheritance from parent cells. This article deals with the molecular structure of chromosomes and with the replication of chromosomes in preparation for cell division.
The primary property of chromosomes, namely the carrying of genes in DNA molecules, is the same in the chromosomes of prokaryotes and eukaryotes. However, prokaryotic and eukaryotic chromosomes differ in both structure and mode of replication in several fundamental ways, and it is convenient to discuss the two types of chromosomes separately. In spite of the differences, the principles of structure and replication already uncovered for prokaryotic chromosomes serve as a basis from which to analyse the much larger and structurally complex chromosomes of eukaryotes.
2. Term Paper on the Prokaryotic Chromosomes:
ADVERTISEMENTS:
A bacterium contains only one chromosome, which encodes all of the genes needed for the life of the cell. Some bacteria contain a very small molecule of DNA called a plasmid, which encodes only a few genes. Some plasmids are valuable to the cell because they confer antibiotic resistance; however, in general, plasmids are dispensable.
Chromosome Size:
Several methods have been developed to measure either the length or molecular weight of the DNA in an intact prokaryotic chromosome. Each of these methods requires chemical extraction of unbroken DNA from cells. Extracting unbroken DNA is difficult because thin molecules easily break by mechanical scission or shear and DNases are present that degrade DNA.
The DNA double helix is a stiff molecule that is easily broken by shearing forces created during stirring, pouring, or pipetting of a DNA solution longer the DNA molecule, the greater the probability of shearing. With simple precautions molecules with molecular weights as large as 108 (150,000 bp) can be isolated without shearing. However, enzymatic breakage can be inflicted by DNases that are activated when a cell is lysed. Because of their small size the DNA molecules of plasmids are the easiest to isolate.
Early estimates of the size of bacterial DNA were low because of shearing and enzyme degradation during preparation. These problems have been solved, and we now know that the sizes of DNA molecules of chromosomes in various bacterial species cover a 15-fold range from 750,000 by to a little over 10,000,000 bp.
Table 8.1 gives the sizes of DNA molecules, measured by various methods, for some bacterial species, a bacteriophage, and a plasmid. Why the DNA molecules of some species of bacteria are so much larger than others is not fully understood. Some bacteria have more genes, giving that cell greater functional versatility. However, it is unlikely that differences in gene number account for more than a part of the differences in DNA amounts.
 One of the first accurate estimates of the molecular weight of DNA in the chromosome in E. coli was made by autoradiography. Cells were grown for two generations with 3H-thymidine to label chromosomes throughout their lengths; then the DNA was extracted very carefully and spread out on a surface for autoradiography. From measurements of the length of autoradiographic images, the DNA was estimated to be at least 1100 μm long, which corresponds to 3,300,000 by or 3300 kilobase pairs (kbp).
One of the first accurate estimates of the molecular weight of DNA in the chromosome in E. coli was made by autoradiography. Cells were grown for two generations with 3H-thymidine to label chromosomes throughout their lengths; then the DNA was extracted very carefully and spread out on a surface for autoradiography. From measurements of the length of autoradiographic images, the DNA was estimated to be at least 1100 μm long, which corresponds to 3,300,000 by or 3300 kilobase pairs (kbp).
For inter conversion of units of size for DNA it is only necessary to remember the 1-2-3 rule:
ADVERTISEMENTS:
1 μm of DNA duplex = 2 × 106 molecular weight = 3 kbp
Three kbp have an absolute mass of 3.1 × 10-6 picograms. One picogram (pg) equals 10-12g. More recent measurements by several methods have shown that the chromosome of E. coli is about 4700 kbp.
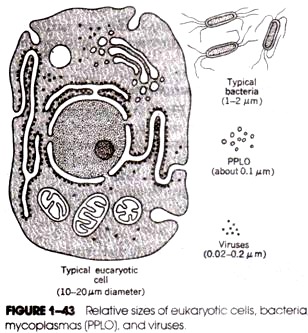
The sizes of smaller molecules, such as those of bacteriophages and plasmids, can often be measured directly by electron microscopy. In general, however, the large molecules of bacterial chromosomes cannot be measured by electron microscopy because the field of view in an electron microscope is small compared with the length of the DNA. An exception is the DNA molecule in the wall-less bacteria, the Mycoplasma.
These are the smallest kinds of bacteria, some measuring only 0.3 μm in diameter. The intact DNA molecule extracted from Mycoplasma hominis measured by electron microscopy is about 250 Am long, which corresponds to 750 kbp. The mycoplasma contains the smallest cell chromosomes known. Assuming as a rough estimate 1000 by per gene (coding sequence plus regulatory sequences) and assuming that, all of the DNA is used for genes, the mycoplasma have an upper limit of 750 genes. About 350 kinds of protein molecules have been identified in mycoplasma.
ADVERTISEMENTS:
Some proteins may have escaped detection, so the number of protein-encoding genes may be somewhat higher than 350. In addition, there are genes encoding rRNA and tRNA molecules. Therefore, the true number of genes in mycoplasma is somewhere between about 400 and 750. Because of its relatively small size the DNA molecule in mycoplasma might be the first cellular chromosome for which the entire nucleotide sequence will be determined. From the sequence virtually every gene and its product will be identifiable.
The chromosome in E. coli is over five times larger than the one in mycoplasma (4700 vs 750 kbp) and could encode 4700 genes. Genetic studies indicate that the number may be 1500 to 2000. Compare this number to the 50,000 to 100,000 different genes estimated to be present in a human cell.
Circularity of Prokaryotic Chromosomes:
ADVERTISEMENTS:
The DNA molecule of all bacterial chromosomes and plasmids is a closed circle. Circularity was first suggested by genetic maps constructed for the chromosome of E. coli. One could explain the positional relationship among various genes in the chromosome by assuming that the chromosome was continuous, that is, was a circle. But there were other explanations for the positional relationships among genes. Proof of circularity came from physical studies of bacterial chromosomes.
The first physical evidence of circularity was obtained from the autoradiographic study of E. coli chromosomes, shown in Figure 8.2. This particular chromosome is in an intermediate stage of replication as reflected by the doubling of part of the circle.
DNA molecules isolated from a variety of plasmids and prokaryotic viruses have been shown by electron microscopy to be closed circles. The chromosome of M. hominis (750 kbp) is the largest chromosome shown to be circular by electron microscopy. The significance of circularity of chromosomes is not known, but it is important in chromosome replication.
Structural Organisation of Bacterial Chromosomes:
Bacterial cells measure no more than a few micrometers in their maximum dimension but contain a DNA molecule hundreds of micrometers in length. E. coli is a cylinder about one μm in diameter and two μm long and contains a circular DNA molecule about 1560 μm long. The DNA molecule fits into the bacterial cell by being compactly folded in a way that does not interfere with transcription or replication.
The folding is obvious in an electron micrograph of a bacterium. The DNA, complexed with protein molecules, forms a dense fibrous mass, the nucleoid. Removal of the tough cell wall by digestion with the enzyme lysozyme allows the cell to be gently lysed. The nucleoid can then be separated from the cytoplasm and studied by electron microscopy.
The DNA is held in the compacted state in the nucleoid by several kinds of proteins. Some of these proteins aggregate to form particles that in turn bind to DNA by means of ionic bonds between positively charged side groups of amino acids, particularly the amino group (—NH3+)of arginine, and the negatively charged phosphate groups of the DNA.
The DNA coils around successive protein aggregates, producing a beaded string in which the extended DNA molecule is drawn into a shortened configuration. The beaded string is itself drawn into a more compacted state, probably by complexing with other proteins, to form the nucleoid. The beaded string, which in its fully extended state would be several hundred micrometers in length, is organised into loops called domains, with all the loops anchored at their two ends to one central core.
ADVERTISEMENTS:
This organisation becomes clear when certain of the proteins, including those that aggregate to form the beads, are removed from an isolated nucleoid. Loops of DNA, each representing a domain containing many genes, extend out from the central protein core of the nucleoid. Because the chromosome is a continuous, circular DNA molecule, one loop must be joined to the next within the central core. This scheme of the organisation of the bacterial chromosome in a nucleoid is shown in the drawing in Figure 8.6.
The organisation of DNA molecules into loops or domains is thought to be important in transcription. The DNA double helix within a loop is under wound. To visualise what this looks like an analogy with a piece of string is useful.
Some kinds of string consist of two threads wound around each other to form a double helix although the demonstration will, work whatever the number of threads. When the two ends of the string are held and then twisted in a direction intended to unwind the two threads, the string coils upon itself, that is, becomes supercoiled.
If one end of the string is released, the string is free to rotate on its own axis and the supercoiling is released. Supercoiling induced by under-winding a double helix is called negative supercoiling. Supercoiling can also be produced by over-winding a double helix; such supercoiling is said to be positive.
Negative supercoiling is shown for the circular duplex molecule of a bacterial virus in Figure 8.7. Because the molecule has no free ends the supercoiling is stable. If a break is introduced into the phosphodiester backbone of the duplex, the bond opposite the break (in the other chain) will act as a swivel and allow the supercoiling to be released.
ADVERTISEMENTS:
The portion of the bacterial DNA double helix in a loop is under-wound and is therefore negatively supercoiled. The DNA is prevented from rotating and releasing the supercoiling because the two ends of a loop are immobilised by attachment to proteins in the central core of the nucleoid.
Twisting of the DNA double helix to produce negative supercoiling is catalysed by an enzyme called DNA gyrase. The proteins that bind to DNA to form the beaded structure of the chromosome then stabilise the supercoiled configuration of individual loops.
Negative supercoiling tends to destabilise the base pair binding between the two chains in the DNA double helix, allowing the chains to separate in localised regions. This separation is believed to facilitate transcription by making the coding strand of the DNA double helix more accessible to RNA polymerase.
Replication of Bacterial Chromosomes:
Replication of the chromosome is a central event in the reproduction of a cell. During cell division the two identical DNA molecules produced by replication are distributed to the two new daughter cells. After cell division the DNA molecule in each daughter cell begins to replicate in preparation for the next cell division. What triggers DNA replication is not known but it is coordinated with the growth of the cell.
Initiation of DNA Replication:
Initiation of replication in E. coli begins at a special region of the DNA molecule called the origin or ori. The origin has been defined by mutational analysis as a 245-bp stretch of the chromosome. That is, mutations in which individual pairs are changed within the 245-bp region decrease the efficiency of initiation of replication. Base changes in adjacent regions are without effect on initiation.
Ori contains several short, repeated sequences that are probably recognised by proteins required for initiating DNA replication. Three of these proteins are encoded by genes designated DnaA, DnaB, and DnaC. Their roles in initiation are only partially understood.
The first step in initiation is binding of the protein encoded by DnaA to the origin, and hence the DnaA protein has been named the initiator protein. What controls the timing of binding (and therefore the timing of initiation) or how it causes initiation is not known. The function of the protein encoded by the gene DnaC is also essential, but precisely what it does is less clear.
The gene DnaB encodes a protein that is required to promote unwinding of the double helix (a helicase). Unwinding gives a special RNA polymerase called primase access to the two strands as templates for replication. The RNA primase synthesizes a short RNA primer, and as shown in Figure 8.9. DNA polymerase then takes over from RNA primase, continuing to extend the new chain with deoxynucleotides.
The unwinding of the double helix promoted by the DnaB helicase causes rotation of the double helix. This rotation is accomplished without inducing supercoiling by the breaking of one chain (nicking), which allows the bond opposite the nick in the other chain to act as a swivel that releases the torsion created by unwinding.
Swiveling is done by an enzyme called topoisomerase, which continuously introduces and reseals nicks ahead of replicating templates to permit progressive unwinding as replication proceeds. Another protein binds to single-stranded DNA and is called single-stranded binding protein or SSB. This protein holds the two strands of DNA in the single-stranded state ahead of DNA polymerase as it moves along the single-stranded template.
Once replication has been initiated with an RNA primer, DNA polymerase works continuously on one template strand but works discontinuously on the opposite strand by the backfilling mechanism. The first region of the chromosome to replicate is the origin, yielding origins in two partial daughter chromosomes.
Whatever condition in the cell caused the parental origin to be activated, it has dissipated, since the two new origins are not used to initiate new rounds of replication until the cell has grown, divided, and again met conditions needed for initiation. A molecular definition of the requirements for initiation of DNA replication and the molecular relation of initiation to cell growth remains one of the most important unsolved problems in cell biology.
Bidirectional Replication:
Figure 8.9 shows the synthesis of an initiation RNA primer on each strand of the newly opened DNA double helix in the origin giving rise to bidirectional replication. By the rules of polarity of the chains in a double helix, two RNA primers are synthesized in a 5’→ 3′ direction, one on each parental template, with nucleotides being added at the end with the free 3’OH group. DNA polymerase replaces primase and extends the RNA chain with deoxynucleotides in the same direction as the primate.
The continuous extension of RNA initiation primers and then DNA chains leaves a single-stranded parental chain immediately opposite. These are converted to double-helix DNA by additional RNA primers made as the parental double helix unwinds. These RNA primers are extended as DNA chains, the RNA primers are removed by exonuclease, and the individual segments of DNA are joined by DNA ligase to complete the double helix.
Because filling in replication with Okazaki fragments necessarily comes after the synthesis of the continuous chain, called the lagging strand, the backfilled strand is called the lagging strand. The overall process as bidirectional replication of the chromosomal DNA, with two replication forks moving in opposite directions from the origin. Autoradiography provided direct visualisation of bidirectional replication in the following experiment.
An auxotrophic mutant of E. coli that cannot synthesize arginine or thymine was grown in the presence of arginine and thymine and was then transferred to nutrient medium lacking arginine and incubated for several hours. Without arginine there is no net synthesis of proteins. In the absence of protein synthesis DNA replication cannot be initiated, and all the bacteria become arrested in the cell cycle at a point shortly before initiation of replication.
Addition of arginine to the medium allows protein synthesis to resume, following which all the bacteria begin to replicate their chromosomes at nearly the same time, that is, the millions of cells in the culture have been synchronised with respect to initiation of DNA replication.
At the moment of addition of arginine to the arginine-starved culture 3H-thymine of low specific activity was also added to the culture. (Low specific activity means that a small fraction of the thymine molecules possess a tritium atom.) When the cells begin replication, the incorporation of the low-specific-activity 3Hthymine results in a low level of labeling of the newly synthesized DNA chains, and a weak autoradiographic pattern will be produced by the DNA.
A minute or two after replication had begun,3H-thymidine of high specific activity (roughly one 3H per thymidine) was added to the culture. Two things are important about the 3H thymidine. First, thymidine is a deoxynucleoside and is incorporated into DNA preferentially to thymine, the free pyrimidine base. Second, the addition of 3H-thymidine at high specific activity results in an immediate shift to incorporation of an intense level of radioactivity at replication forks and a resulting intense autoradiograph from the DNA. After a minute or two of incorporation of 3H-thymidine, the bacteria were cooled to stop further DNA replication, treated with lysozyme to digest the cell wall, and a drop of the culture was lysed with a drop of ionic detergent on a microscope slide. The detergent stripped the proteins from DNA.
The drop of culture detergent mixture was spread, which caused DNA to be extended out onto the microscope slide. Autoradiography then revealed the pattern of radioactive regions in DNA molecules. Each molecule produced a central light autoradiograph for the two partial daughter segments (3H- thymine incorporation).
The light pattern represented the first DNA to replicate after initiation. The intense autoradiographic patterns represented subsequent synthesis and showed that replication had proceeded bi-directionally from the origin (the origin is at the center of the light autoradiograph). The two partial daughter molecules were joined at the replication forks at the outer extremities of the autoradiographic patterns. The un-replicated, parental DNA ahead of the replication fork was not radioactive and hence not seen in the autoradiograph.
Autoradiography shows the overall pattern of replication but does not have the resolving power to reveal the finer molecular events such as backfilling with Okazaki fragments, which occurs within the first micrometer behind the replication fork.
During replication, proteins that bind to DNA to produce the folding pattern of the chromosome, as well as proteins involved in transcription (RNA polymerase, repressor molecules, etc.) are probably transiently displaced from the DNA as the replication forks move through it. New proteins that make up the replication enzymatic machinery as well as single-stranded binding protein (SSB) bind to the DNA in the immediate region of the replication fork and migrate with it.
The SSB binds only to single-stranded DNA and stabilises the lagging parental strand prior to its replication. Behind the fork the newly replicated segments of daughter double helices become folded and packed by binding with packing proteins and may resume transcription with binding of RNA polymerase and other transcription factors.
All of these events-releases of packing proteins, displacement of transcription factors, opening the double helix, and replication-occur extremely rapidly. At its maximum rate a replication fork moves through 980 by of DNA per second.
Termination of Replication:
The two replication forks, traveling in opposite directions, eventually meet (in 40 minutes at the maximum rate of DNA replication) on the opposite side of the circular chromosome. Figure 8.10. shows an autoradiograph of two replication forks approaching each other less than a minute before termination. Termination apparently does not occur at a specific deoxynucleotide sequence.
Instead, in the termination region the rate of fork travel is enormously slowed (by an unknown means), and forks always meet in roughly the same region of the chromosome. The two replication forks abolish each other on meeting.
At this point the two circular daughter molecules are complete, but are still inter-looped with each other in concatenated fashion. The final step in replication is separation of the loops. Separation is catalysed by a topoisomerase, which creates a transient double stranded break in one of the daughter double helices, allowing the other to slip through, and then reseals the double helix.
3. Term Paper on the Eukaryotic Chromosomes:
The structure and replication of eukaryotic genomes are more complicated than in prokaryotes.
Although there are important similarities between the two groups of organisms, there are also major differences:
1. In all prokaryotic organisms the entire necessary set of genes (genome) is carried in one chromosome. In contrast, almost all eukaryotic species distribute their genome over a number of chromosomes, ranging up to hundreds in some species.
2. The aggregate molecular weight of the DNA in a haploid set of chromosomes in a eukaryotic cell is a few times to tens of thousands of times greater than the molecular weight of the DNA in the largest prokaryotic chromosome known. The greater amount of DNA underlies the greater genetic complexity of eukaryotes, although much of the DNA in most eukaryotes has no known function.
3. The chromosome in a prokaryotic cell is always a covalently closed, circular DNA molecule. Each nuclear chromosome in a eukaryotic cell contains a single, linear molecule.
Chromosome Numbers in Eukaryotes:
The smallest haploid chromosomal complement in a eukaryote occurs in a species of ant-the number is one. The plant Haplopapus has two chromosomes, and all of the thousands of other eukaryotes examined so far possess more than two chromosomes. Table 8.2 lists the haploid chromosome numbers for a variety of eukaryotes, and Figure 8.11 shows mitotic chromosomes of several organisms.
The chromosomal numbers show no regular pattern within groups of plants or animals. Different species of mammals have numbers that range from four in the Indian deer to 23 in the human to 39 in the dog. Mammalian species with fewer chromosomes have larger chromosomes.
Some protozoa have as few as five chromosomes (e.g., Tetrahymena); others have hundreds (Amoeba proteus). Large differences occur among plants, algae, amphibians, fishes, reptiles, and so forth, although in most species the chromosome number is less than 100. The significance of such divergent chromosome numbers, which occur even among closely related species, is not known.
Amount of DNA in Eukaryotes:
The total amount of DNA in a haploid set of chromosomes in a species is called the genomic value or C- value of the species. The C- values for eukaryotes exceed the amount of DNA in the largest prokaryotic chromosomes by a few times (yeast) to several million times (some salamanders). This greater amount of DNA underlies the greater genetic complexity of eukaryotic species.
C-values differ greatly among eukaryotes. For example, the C-value for the human is 3.1 pg., for the lungfish 50.0 pg., and for Amphiuma means 100 pg. Even Amoeba proteus has many times more DNA than a human. It might be assumed that more DNA in the genome means a greater number of different genes. However, it can hardly be supposed that salamanders have more kinds of genes than humans. The resolution of this apparent contradiction between DNA amount and expected gene number, called the C-value paradox.
The Size of DNA Molecules in Eukaryotes:
The amount of DNA per chromosome differs over a wide range in eukaryotes. The budding yeast has the smallest chromosomes discovered so far. The C-value of 1.5 × 10′ base pairs of DNA is distributed among 17 chromosomes in a haploid set.
The sizes of the DNA molecules in these chromosomes have been determined partly by electrophoretic migration in a gel. The smallest chromosome contains a DNA molecule of 260,000 base pairs (87 μm), or about 1/16 as long as the chromosome in E. coli. The others extend upwards in size with the largest one well over one million base pairs (over 333 Am in length).
There is no conceptual difficulty in accepting the length measurements for yeast chromosomes; they are all considerably smaller than the DNA molecule in E. coli. Other eukaryotes possess much more DNA per chromosome, in some the equivalent of DNA molecules several meters long. Such long molecules seem improbable, and for years it was debated whether a eukaryotic chromosome contains a single, long DNA molecule or multiple short ones.
The most direct way to determine whether the DNA in a chromosome is present in one or many molecules is to purify the DNA and measure its size directly by some physical technique. This has been done for the relatively short molecules in yeast, but is virtually impossible for much longer molecules because of the ease with which they are broken, especially during purification. Indirect methods for determining molecular length had to be devised as described below.
The lengths of the DNA molecules in chromosomes of various species cover a wide range. Table 8.3 gives the average length of DNA molecules for some eukaryotic species. Why organisms package their genes in such vastly different sizes of DNA molecules is not known.
Length of DNA Molecules in Drosophila:
One method for measuring DNA length exploits an elastic property of DNA. Molecules of DNA free in solution coil randomly, much as a long piece of string coils when immersed in water. When a DNA molecule is pulled into a straight configuration and then released, it recoils back into a random configuration. The time taken for a straight molecule to relax is proportional to its length. Measuring the relaxation time provides an indirect but accurate measure of molecular length.
A great advantage of the method is that the DNA does not need to be purified to be measured, which eliminates manipulations of DNA that inevitably break it. Cells are lysed with an ionic detergent, which strips all the proteins from DNA. The elasticity of DNA is measured directly in the detergent solution.
When applied to cultured cells of Drosophila, the largest DNA molecule gives a molecular weight of 41 × 109, which corresponds to a length of 20.5 mm. The total amount of DNA in the largest chromosome was previously known from optical measurements to be 41 × 109, and the elasticity measurement proves that all in this DNA is contained in a single molecule.
Length of DNA Molecules in Mammalian Cells:
Estimates of molecular length can be obtained for mammalian chromosomes with the autoradiographic technique previously described for visualising the DNA of E. coli. Cultured human cells are first grown for at least one cell cycle with 3H-thymidine, which completely labels the DNA molecules throughout their lengths. The cells are lysed in a drop of dilute ionic detergent on a microscope slide.
Then the drop is gently spread over the slide, which draws the DNA into a long, straight form. Subsequent autoradiography shows the presence of tracks of silver grains several centimeters long, reflecting the presence of DNA molecules of such length.
A human diploid cell contains 6.2 pg. of DNA and 46 chromosomes, which means on average an equivalent of 4 cm of DNA per chromosome. The autoradiographic visualisation shows that all in the DNA in a chromosome is contained in a single molecule.
Length of DNA Molecules in Salamanders:
The genomes and chromosomes of salamanders are among the largest known. Only the African lungfish and a few species of lilies have comparable amounts of DNA per genome and per chromosome. A haploid set of 12 chromosomes in the salamander Amphiuma means (misnamed the Congo eel) contains 100 pg of DNA. Therefore, on the average each chromosome contains the equivalent of 2.7 meters of DNA. According to experimental evidence, some of which is cited below, all 2.7 meters of DNA are present in a single, extremely long molecule.
Most studies of DNA in salamanders have been done on chromosomes in meiosis. In salamanders the formation of oocytes lasts many months. During this time the chromosomes are arrested in the prophase stage of the first meiotic division. Because the chromosomes are partially condensed, they are easily seen by light microscopy.
A single lampbrush chromosome actually contains two homologous chromosomes that previously were tightly joined in meiotic pairing and subsequently have largely separated from each other. The chromosomes are now in the diplotene stage of the first meiotic prophase.
The two homologues have separated except at several joined points called chiasmata (sing. = chiasma). Each chiasma is the cytological manifestation of a crossover event that took place earlier in prophase. Each homologue underwent duplication (DNA replication) in the premeiotic interphase and consists of two chromatids. Crossing over has occurred between a chromatid in one homologue and a chromatid in the other, which holds the chromosomes together at chiasmata.
The fuzzy appearance of the two homologues in a lampbrush chromosome is the result of threads that loop out from its two chromatids. The loops occur in pairs-wherever a loop occurs in one chromatid, it is matched by a loop of the same size in the other chromatid. The loops are made up of DNA with attached ribonucleoproteins and are sites of transcription.
The axis of the chromosome consists of a string of fine chromatin granules called chromomeres. One or more pairs of loops extend from each chromomere. In this arrangement each chromatid consists of a string of chromomeres with attached loops. Since the two chromatids are tightly opposed to each other, the chromosome axis is probably formed by fused pairs of chromomeres.
The molecular structure of chromomeres and loops was deciphered by a combination of experiments. First, removal of proteins and RNA from lampbrush chromosomes with enzymes does not cause breakage of the chromosome. Treatment with DNase causes fragmentation. This suggests that the linear continuity of the chromosome is not dependent on protein or RNA but does depend on continuous intact DNA.
When protein and RNA are stripped from loops, only a DNA thread is left. Measurements of the diameter of the thread seen in loops by electron microscopy show that the thread is a single DNA double helix. A comparison of the rates of breakage with DNase of loops versus the main axis of the chromosome shows that each loop contains one DNA duplex whereas the main axis must contain two DNA double helices, one for each chromatid.
These and other observations showed that each chromatid contains a single DNA double helix that extends through the entire length of the chromatid. Coiling and folding of the DNA gives rise to the successive chromomeres. Successive chromomeres are connected by segments of the DNA molecule that loop out from one chromomere and continue into the next. In short, one DNA molecule traverses the entire length of a chromatid. Proteins are responsible for folding the DNA in a specific pattern of successive chromomeres and loops.
The partial meiotic condensation of lampbrush chromosomes makes visible a structural organisation along the chromosome that may be present in interphase chromosomes of eukaryotes in general, although probably in a looser form. This presumed structural organisation is not discernible because the chromosomes are more extended and intertwined with one another within the nucleus.
Direct support for this view is provided by polytene chromosomes. Polytene chromosomes are true interphase chromosomes. Their linear organisation is visible because many identical copies of the DNA double helix, with attached proteins, are held together in exact parallel register, producing a clearly discernible chromosome. Each band of a polytene chromosome may correspond to a chromomeric granule and its loop.
Organisation of Deoxynucleotide Sequences in Eukaryotic DNA:
The DNA molecule of a prokaryotic chromosome is largely occupied by a succession of genes. A gene is defined here as a coding sequence and sequences upstream and downstream that are concerned with controlling the expression of the coding sequence. No genes have yet been found in some segments of the chromosome in E. coli, but most of the DNA consists of identifiable genes.
Eukaryotic DNA molecules are usually larger than prokaryotic molecules and possess a number of features not observed in prokaryotes. Three distinguishing features were mentioned to introduce eukaryotic chromosomes.
Additional characteristics are the following:
1. The DNA of most eukaryotic genomes contains non-gene sequences that are repeated many times (repetitive DNA).
2. A few kinds of eukaryotic genes are present in multiple copies.
3. Eukaryotic DNA contains imperfect, apparently nonfunctional copies of some genes called pseudo-genes.
4. The linear DNA molecules in eukaryotic chromosomes terminate in special sequences called telomeres, except for mitochondrial and chloroplast chromosomes, which are usually circular and therefore have no ends.
5. Eukaryotic chromosomes possess centromeres, structures by which chromosomes are distributed at mitosis and meiosis. The centromere has its basis in a specific sequence in the DNA molecule.
6. Prokaryotic DNA molecules contain a single origin of replication. The linear molecules in eukaryotes have many origins dispersed along their lengths.
7. Prokaryotic chromosomes are complexed with DNA-binding proteins that package the DNA molecule in successive units. Eukaryotic DNA is bound to a special class of proteins, the histones, forming repeating units called nucleosomes.
8. Most genes in eukaryotes are interrupted at a number of sites by introns. With a few exceptions introns are absent from prokaryotic genes.
9. Much of the DNA in most eukaryotic genomes forms long spacers between successive genes or groups of genes in a molecule. Spacers between genes in prokaryotes are very short.
Repetitive DNA Sequences:
All eukaryotes contain repetitive sequences. A small part of the repetitive sequences represents a few kinds of genes that are present in multiple copies, but most of the repetitive sequences have no known function, genetic or otherwise.
The absolute amount of repetitive sequences tends to be greater in organisms with larger genomes. In yeast cells only a few percent of the total DNA consists of repetitive sequences. Typically in mammals 30 percent to 40 percent of the DNA is made up of repetitive sequences belonging to several families.
The remaining 60 percent to 70 percent is made up of sequences that occur only once per haploid genome. These are called unique sequences and contain most of the genes as well as a large amount of DNA of unknown function.
Within a family of repetitive sequences all the sequences are identical or nearly identical with one another and differ from sequences in any other family. Families are classified according to the number of copies of the repeat. Families containing two to 100 copies are slightly repetitive.
Families with 100 to several thousand copies are middle repetitive. Families with several thousand to several million repetitive sequences are called highly repetitive. Some of the families of slightly or middle repetitive DNA are, in fact, multiple copies of certain genes.
For example, genes encoding actin are repeated several times in most eukaryotes and therefore make a family of slightly repetitive sequences. Genes encoding histone proteins are usually present in a few hundred copies and represent middle repetitive DNA. However, the functions, if any, of most families of middle repetitive sequences are unknown.
An example of a highly repetitive sequence occurs in the centromeric region of mouse chromosomes. This sequence is about 300 base pairs long, is repeated about 106 times per genome, and accounts for 10 percent of the DNA in the mouse genome. It occurs in blocks of thousands of copies in tandem arrays. Its location in the centromeric region of chromosomes implies a centromere-related function, but its function is not known.
Blocks of highly repetitive sequences are packaged in a tightly condensed heterochromatin, which is in accord with the observed absence of transcription. Blocks of highly repetitive sequences often differ in overall base composition from the rest of the DNA in the genome. If the repetitive DNA has a higher-than-average content of AT base pairs compared with the rest of the DNA, it has a lower-than-average density in a CsCI solution.
GCrich repetitive DNA has a higher average density. When total mouse DNA is cut into segments of a few hundred base pairs and centrifuged to equilibrium in a solution of CsCI, the highly repetitive DNA forms a small band adjacent to the main band of DNA because it has a higher average AT content and therefore a lower density than the main DNA. This, in fact, was how highly repetitive DNA was first discovered, and it is sometimes referred to as satellite DNA because it forms a satellite peak in a density profile.
The individual copies of sequences in families of all three classes of repetitive sequences (slightly, middle, and highly repetitive) can occur in blocks as just described for the family of highly repetitive sequences in the mouse genome, or the individual copies can be dispersed apparently randomly among genes throughout the genome. Middle repetitive DNA is also usually distributed in the genome in between genes.
An example of a highly repetitive sequence that is dispersed is the Alu family in the human genome. The repetitive sequence in the Alu family is about 300 base pairs long, more than 900,000 copies are present, and they account for five percent of human DNA.
One or more copies have been found not far from all genes isolated. At least some Alu sequences are transcribed, forming part of the primary transcript of some genes. The Alu segments of primary transcripts are removed and destroyed in processing of the transcript in the formation of an mRNA molecule. Like all repetitive sequences the function of the Alu sequences or their transcripts is not known. Closely related species sometimes have greatly differing amounts of repetitive DNA.
For example, the genome of Drosophila melanogaster contains about 15 percent of highly repetitive sequences, D. virilis has 40 percent, and D. cyrtoloma has 60 percent. In one species of the kangaroo rat 60 percent of the DNA is repetitive; in a closely related species less than ten percent is repetitive, and the total amount of DNA in the genome is correspondingly less.
Telomeric DNA Sequences:
The linear DNA molecules of eukaryotes have special sequences at their ends called telomeric sequences. The sequence of telomeric DNA has been determined in a number of lower eukaryotes. Although there are differences, all telomeric sequences conform to a common pattern. In Tetrahymena and Paramecium, telomeres consist of up to 70 repeats of the sequence 5’CCCCAA3′.
In a different group of ciliates (the hypotrichs) telomeres consist of several repeats of the sequence 5’CCCCAAAA3′. In a flagellated protozoan the repeated sequence in telomeres is 5’CCCTAA3′, in yeast it is more variable, consisting of 5’CCACA3′ and slight variations of the sequence. The telomeric sequences are important in conferring stability on linear DNA molecules, since experimentally introduced linear DNA molecules that lack them are degraded.
Autonomously Replicating Sequences:
An autonomously replicating sequence or ARS is so named because it confers autonomous replicative capacity on a circular plasmid in yeast cells. It is comparable in function to the on sequence in a prokaryotic chromosome. The ARS in the yeast plasmid can be deleted, and a segment of foreign DNA inserted in its place to test for ARS activity of the foreign DNA in yeast cells.
Segments with ARS activity have been found in this way in chromosomal DNA of yeast and other organisms including human DNA, and these may be sequences defining origins of replication in their normal locations. The ARSs in yeast chromosomes differ somewhat from one another in sequence, but all those isolated so far have an 11- base-pair, AT-rich sequence with at least 200 base pairs flanking each side, which are necessary for normal initiation of replication. An ARS presumably is recognised by a specific protein that triggers the initiation of DNA replication, but such a protein has not yet been identified in eukaryotes.
Centromeric Sequences:
The proper distribution of chromosomes in eukaryotes during mitosis and meiosis is achieved by attachment of chromosomes to microtubules in the spindle apparatus in the dividing cell. The attachment in most species occurs at a region in the chromosome called the centromere. The centromere is also the point at which the two chromatids remain attached to each other in metaphase.
Separation of the two chromatids at the centromere marks the transition from metaphase to anaphase; from that instant on, the chromatids are called daughter chromosomes. The attachment of microtubules occurs at a part of the centromere called the kinetochore, which in the electron microscope looks like a plate on each chromatid. The centromere is defined by a sequence of deoxynucleotides. Special proteins probably bind to the centromeric sequence and give the centromere its functional properties and form the kinetochore.
Centromeric sequences from several chromosomes (CEN sequences) have been studied closely in yeast. They are quite similar but not identical in sequence, consisting of 80 to 90 base pairs of 93 to 94 percent AT base pairs next to a highly conserved 11-base-pair sequence. A CEN sequence can be removed from a chromosome and reinserted in the opposite orientation, and it works just as efficiently. CEN sequences, with slightly different composition, can be interchanged among yeast chromosomes without any loss of function.
Pseudogenes:
Pseudogenes are nonfunctional, imperfect copies of normal genes. They occur commonly in higher eukaryotes such as mammals, but only a few have been found in lower eukaryotes such as yeast and Drosophila. In mammals pseudogenes account for as much as one percent of the total DNA. Formation of a pseudogene is a rare event in evolution, but once formed a pseudogene remains even though it is nonfunctional.
For example, pseudogene Jn is a defective gene for β-globin of hemoglobin in mammals. A detailed evolutionary history of pseudogene Ψƞ has been constructed by the study of the globin gene family in various animals, and it is clear that pseudogene Ψƞ arose more than 140 million years ago, long before the evolution of modern mammals.
Pseudogenes arise by two mechanisms. Some, such as Ψƞ originated by chance duplication of a normal functional gene. Duplication of a gene can occur by unequal crossing over during meiosis such that one chromatid receives both copies of a gene and the other chromatid receives no copy.
Gene duplication can also occur through a mistake in DNA replication. A segment of DNA is replicated twice, and the extra copy becomes integrated into a chromosome by a poorly understood mechanism of breakage and rejoining. The result is the presence of two functional copies of a gene.
Duplication of a gene may be followed sometime later in evolution by mutation of one of the copies into a nonfunctional form, creating a pseudogene. Defects observed in pseudogenes include abnormal initiation and termination codons, missense, nonsense, and frameshift mutations, altered promoters, and abnormal splicing in transcripts.
In the second mechanism pseudogenes are formed from mRNA molecules. Certain RNA viruses, called retroviruses, carry a gene encoding an enzyme that can synthesize double-stranded DNA using a single stranded RNA chain as a template instead of DNA. The enzyme is called reverse transcriptase.
The resulting segment of DNA double helix, called cDNA, may become integrated into chromosomal DNA by breakage and resealing. The integrated cDNA, being a copy of mRNA, lacks a promoter and other sequences necessary for transcription and hence cannot be transcribed. Pseudogenes formed from mRNA lack the introns present in most normal genes, and are therefore easily distinguished from their normal, intron-containing counterparts.
Sequence analysis suggests that some families of dispersed repetitive DNA sequences are, in fact, pseudogenes of tRNA molecules, snRNA molecules, or other small non-mRNA molecules that are plentiful in the cell and therefore more likely to have been used by chance to make a cDNA. Thousands of nonfunctional, pseudogene copies of snRNA genes are found in mammals. The human genome contains about 20 copies of the gene encoding actin; many of these are nonfunctional pseudogenes.
Junk DNA:
Pseudogenes, other dispersed repetitive sequences not clearly identified as pseudogenes, and perhaps other non-repetitive sequences with no known function may all be defective relics of normal genes. For that reason they are sometimes called junk DNA.
If such DNA sequences contribute nothing to the functioning of a cell, that is, are genetically neutral, it is not clear why they are not in the course of time lost through chance deletion. The cost of carrying junk DNA, for example, in the expense to the cell of its replication, may be so trivial that organisms containing it do not incur a survival disadvantage.
The Essential Sequences of a Chromosome:
Many kinds of sequences have just been listed; some are essential for chromosome structure and function, and some are not.
The minimal sequences in a chromosome are genes, telomeres, ARSs, and a centromere. So far, research has hot identified functions for most repetitive sequences (not including normal gene families) and pseudogenes, and they are presumably nonessential.
The sequences required for chromosome function have been defined experimentally in yeast cells in the following way- a linear piece of DNA containing an ARS and a few genes was combined with a CEN sequence, and telomeres were joined to the ends. The molecule, containing about 104 bp, was introduced into yeast cells. Although the molecule persisted through several cell divisions, it was lost because it was not properly distributed to daughter cells by mitosis.
Apparently, the CEN sequence did not function adequately in such a short DNA molecule. A major difference between this artificially constructed chromosome and a normal yeast chromosome is length. The smallest yeast chromosome contains 2.6 × 105 bp, compared to the 104 by in the artificial chromosome. The artificial chromosome was therefore enlarged to 5 × 104 by using DNA from a bacteriophage.
Bacteriophage DNA is functionless in yeast and was used to act as filler. Increasing size in this way markedly improved the mitotic stability of the artificial chromosome and even allowed it to behave normally in meiosis, but it was still not as stable as the larger normal chromosomes in yeast. A normal yeast chromosome is accidentally lost with a frequency of once per 104 to 105 cell divisions. Smaller chromosomes are lost more frequently than larger ones.
These experiments imply that a DNA molecule, in addition to possessing ARS, telomeres, genes, and a CEN sequence must have some minimum size in order to be distributed properly in mitosis and meiosis. The apparent size requirement is met in yeast almost exclusively by genes, with little or no contribution from repetitive sequences, pseudogenes, or junk DNA.
Total Complexity of DNA Sequences in Genomes:
The total complexity of sequences, as opposed to total DNA, is the sum of all the different sequences in a genome. In prokaryotes total sequence complexity is essentially equivalent to the DNA content of the genome since almost all the DNA consists of single copy, or unique sequences.
In eukaryotes, total sequence complexity is less than the total DNA content because of the presence of repetitive DNA. Repetitive DNA accounts for a significant part of the total DNA but because it is composed of the same sequence repeated over and over, it contributes disproportionally less than unique-sequence DNA to total sequence complexity. For example, about 25 percent of the DNA in the mouse genome is repetitive and 75 percent is unique. Of the 25 percent repetitive DNA, 10 percent is a highly repetitive sequence of 300 by (the satellite DNA), and 15 percent is middle repetitive DNA.
The total DNA content in the mouse genome is about 3 × 109 bp. The unique-sequence DNA (75 percent of total or 2.25 × 109 bp) accounts for virtually 100 percent of the sequence complexity. The highly repetitive sequences accounts for 10 percent of the total DNA but only 0.000013 percent of the total complexity (300 bp/2.25 × 109 bp).
The several families of middle repetitive DNA account altogether for less than one percent of the total sequence complexity in the mouse genome. The sequence complexity of DNA is determined by shearing it into fragments and dissociating the two chains of the double helix by denaturation; then, the rate at which they re-associate is measured as a function of DNA concentration expressed as nucleotides per liter. The greater the complexity of DNA, the more slowly it re-associates. Consider, for example, two DNA molecules each consisting of 1000 bp.
Suppose one of the molecules consists entirely of unique sequences and the other consists of a 100-base-pair sequence repeated identically ten times. Suppose the same number of molecules of 1000 bp is dissolved in a salt solution of the same volume, cut in segments about 100 bp long, denatured, and then allowed to re-nature. The rate of re-association is governed by how often complementary chains encounter each other in solution.
The concentration of chains of particular sequences is ten times higher for the repetitive DNA than for the unique sequence DNA; therefore the repetitive DNA will re-associate much faster than the unique sequence DNA. Likewise, when the total DNA of a genome is cut into lengths of a few hundred by and dissociated, each kind of sequence re-associates at a rate that is proportional to its concentration; highly repetitive sequences re-associate more rapidly, followed by re-association of middle repetitive sequences, and then by re-association of the unique sequences.
Figure 8.18 shows re-association curves for DNAs of T4 bacteriophage, E. coli, and the mouse. Since T4 and E. coli DNA do not contain repetitive sequences, they re-associate as uniform curves reflecting rates expected for genomes of their respective sizes. The genome of E. coli DNA is about 25 times larger than DNA of T4. If both DNAs are prepared at the same concentration (in nucleotides per liter), the DNA of E. coli re-associates more slowly than T4 DNA (because the number of DNA molecules is less).
Mouse DNA shows a more complicated curve, the first part of which reflects re-association of highly repetitive sequences (satellite DNA), followed by several families of intermediate repetitive DNA and finally unique sequence DNA.
A useful way to record re-association events is to plot the initial molar concentration of nucleotides (Co) in the DNA times the time (t) in seconds that re-association has been occurring against UV absorption; a decline in UV absorption occurs as DNA re-associates. Such a plot of initial Co × t (Cot) against UV absorption is shown in Figure 8.18. The UV absorption of totally dissociated DNA at t = 0 is defined as 100 percent. The value of Cot when half the DNA has re-associated, Cot1/2, is used as a measure of the sequence complexity of the DNA in nucleotides. That the Cot1/2, for the E.coli DNA is about 25 times greater than for T4 DNA, and the Cot1/2 for the unique fraction of mouse DNA is about 600 times greater than for E.coli DNA.
The measure of sequence complexity of genomes, or S-value (S for sequence), is in some ways a more useful value than is total DNA since total DNA can differ by wide margins from species to species because of large differences in the amounts of repetitive sequences.
The DNA content of mammals covers a twofold range, but all have about the same complexity, which is about 600 times the complexity of E. coli DNA. This does not mean that mammals possess 600 times more genes than E. coli because most of the DNA sequences in mammals do not encode genes.
The C-Value Paradox:
The C-value paradox mentioned earlier occurred because of wide differences in DNA content among species. Comparing S-values among organisms instead of comparing C-values might be expected to show a closer correspondence. After all, S-values are measures of the amount of unique sequence complexity, and they avoid the wide differences in total DNA that stem from differences in the content of repetitive DNA.
The correspondence is, indeed, closer when S-values are used. All mammals have close to the same structural and functional complexity, and although C-values vary over a two-fold range, they all have very close to the same S-value. However, among eukaryotes a lack of correspondence between S-values is still often observed. For example, salamanders have several times more unique-sequence DNA than do mammals. The solution to this S-value paradox is that a large amount of unique-sequence DNA does not encode genes.
Estimates of Gene Numbers, the S-Value Paradox, and Spacer DNA:
The precise number of genes is known for a few of the simpler viruses, but it is not known for any prokaryotic or eukaryotic cell. Assuming 1000 by for one gene, mycoplasma contains a maximum of 750 genes. The chromosome of E. coli contains 4.7 × 106 bp, enough DNA for 4700 genes, although the number is certainly much less than that. Eukaryotes contain more genes than prokaryotes, but at present only rough estimates are possible.
Gene Number and Spacers in Drosophila:
Drosophila melanogaster contains 1.65 × 108 by of DNA, of which 85 percent or 1.40 × 108 by is unique sequence DNA. Assuming a somewhat larger average size for the coding regions of genes in Drosophila, such as 1200 bp, 1.40 × 108 by could contain 117,000 genes. Estimates based on other grounds are considerably less.
Genetic evidence suggests that one band of a polytene chromosome corresponds to one gene, although some bands may contain more than one gene. There is a little over 5000 bands total in the four polytene chromosomes, so estimates of gene number are 5000 to 7000. At 1200 by per gene there is a 17- to 23-fold excess of unique sequence DNA.
A clue about the disposition of the excess unique sequence DNA is apparent from the study of the polytene chromosomes in Drosophila. On the average, bands in a polytene chromosome contain segments of DNA consisting of about 25,000 by of unique sequence.
Using the estimate of 1200 by for a gene, with one gene per band, about five percent of the DNA in a band is used to encode a gene. Some part of the remaining 95 percent (23,800 bp) can be accounted for by introns and pseudogenes. However, the function of the bulk of the unique-sequence DNA cannot be accounted for, and at present it must simply be regarded as spacer DNA between successive genes.
Gene Number and Spacers in Mammals:
In mammals the amount of unique sequence is sufficient to encode over 1.5 × 106 genes. A number of genetic considerations suggest a number 15 to 30 times less, or between 50,000 and 100,000 genes. Most eukaryotic genes contain introns, sometimes many of them, and some are very large.
Even including introns as parts of genes does not resolve the S-value paradox. The solution is, again, probably the presence of large spacers of unique sequences between successive genes. For example, a 50,000-bp segment of DNA in humans contains the five members in the family of the, β -globin genes and the Ψƞ pseudogene for β -globin.
Each gene consists of about 2/3 intron sequences and 1/3 coding sequences. However, the six genes, including their introns and known regulatory sequences upstream and downstream from the genes, account for only about 20 percent of the 50,000-bp segment. The rest of the DNA, most of which consists of unique sequences, contains no identifiable genes and appears to exist as spacers between members of the globin gene family.
Gene Number and Spacers in Ciliated Protozoa:
In ciliated protozoa it is clear that most of the unique sequence DNA in the eukaryotic chromosomes exists as spacers between genes and has no known function. A ciliate contains a small, diploid micronucleus and a large, DNA-rich macronucleus. The micronucleus does not produce RNA, and it functions during cell mating. The macronucleus makes all the RNA needed for cell maintenance and reproduction.
The major activity of ciliates is growth and division but occasionally two cells mate (conjugate). During conjugation the micronucleus undergoes meiosis, and the cells exchange haploid micronuclei and then separate. Immediately after separation the macronucleus is destroyed, and a new macronucleus is formed from a micronucleus.
In one group of ciliates known as hypotrichs, the first stage in macronuclear development is the formation of polytene chromosomes by multiple rounds of DNA replication. The process takes two days. As soon as they have formed, the polytene chromosomes are cut into short segments, each segment corresponding to a band of a polytene chromosome.
A band is believed to represent a single gene, and the cutting up of the polytene chromosomes is; therefore, a gene-by-gene process. Cutting produces segments of DNA that contain on the average over 40,000 bp. These are rapidly reduced to an average of 2200 bp by cleavage of most of each DNA segment to nucleotides.
Each of the short molecules that is left represents a single gene. Short telomeric sequences are then added to the ends of each gene. Finally, all the genes replicate many times to produce a mature DNA-rich macronucleus in which all the DNA occurs in short molecules, each encoding a single gene.
The mature macronucleus contains about 20,000 different DNA molecules, each present in about 1000 copies. These genes, representing only five percent of the sequence complexity in the chromosomes of the original micronucleus, are transcribed to provide all the mRNA, tRNA, rRNA, and other RNAs needed by the cell.
In summary, the development of the macronucleus consists of excision of all the genes from the chromosomes and destruction of most of the sequences in the chromosomes. Why polytene chromosomes are formed as an intermediate stage and many of the molecular details of how genes are excised and supplied with telomeres are still not known.
However, the unusual manipulations of the genome by these ciliates reveal how gene and nongene sequences are organised in chromosomes. The gene-sized molecules in the macronucleus can be used as hybridisation probes to map the position of the genes in chromosomal DNA using long segments of DNA cloned from micronuclear chromosomes.
Genes are found singly or in small groups along the DNA molecule separated by long spacers of nongene DNA. It is the spacer DNA, accounting for 95 percent of the total deoxynucleotide sequences, that is degraded following the gene-by-gene segmentation of the polytene chromosomes. This extreme elimination of all nonessential sequences to form the functional nucleus presumably provides some advantage to these organisms.
Ciliates may require the many copies of each gene in the macronucleus to support their very large cell size. By elimination of all nonessential sequences, a high copy number for each gene can be achieved with a minimum amount of DNA and a minimum expansion of nuclear size.
The several observations on the DNA of Drosophila, mammals, and ciliates are consistent in suggesting a general model for eukaryotes, in which most of the DNA sequences form spacers of unknown significance between successive genes. Large differences in DNA sequence complexities (S-values) among eukaryotes, such as the high values in salamanders and lilies, are probably due to large differences in the size of spacers between genes.
Gene Families:
Most genes in both prokaryotes and eukaryotes are present in single copies per genome. Although a few kinds of genes exist in multiple copies, in general single copies of genes are sufficient, because expression of genes that encode protein products is amplified at two points. First, many mRNA copies are transcribed from a single gene, and second, each of the many mRNA copies is translated into many copies of the protein.
For most genes these amplifications at the transcriptional and translational levels are sufficient to meet the needs of a cell. Even during differentiation of cells, when a large amount of a gene product may be required, single copies of genes are sufficient. For example, an enormous number of hemoglobin molecules are produced in the formation of a red blood cell. Single copies of genes encoding α- and β-globin are sufficient.
For genes whose final products are RNA molecules, like genes encoding tRNAs and rRNAs, expression is by transcription alone. For these genes, as well as for a few kinds of genes that encode proteins, the rate of production of product cannot be achieved with a single copy of the gene per genome because of limits on rates of transcription (10 to 50 mRNA transcripts per gene per minute and 3 to 20 polypeptides per mRNA molecule per minute).
Almost all eukaryotes have multiple copies of the sequences encoding the four rRNAs. Sequences for three of the rRNAs (28S, 18S, and 5.8S in mammals) are grouped in a single transcription unit of DNA and are transcribed as a single precursor rRNA that is post-transcriptionally processed into three final products.
Transcription units for the three rRNAs are repeated in tandem in the chromosome in many copies. In yeast cells the transcription unit is repeated 100 to 120 times in the arrangement shown in Figure 8.21. Successive transcription units are separated by about 2000 bp. Within the 2000 bp is a gene of about 120 bp that codes for the fourth rRNA, 5S rRNA.
The rest of the 2000 bp is spacer. In other eukaryotes the 5S genes occur elsewhere in the genome as separate tandem arrays. In these eukaryotes the spacers between the transcription units for the three rRNAs contain no coding function, and no part of them is transcribed.
The size of spacers between rRNA transcription units ranges from about 2000 bp in some protists to 50,000 bp in some higher eukaryotes.
There are 450 rRNA transcription units per haploid genome in Xenopus, arranged in tandem. Each is 7600 bp long and separated from adjacent units by un-transcribed spacers of several thousands of base pairs. The transcription of rRNA genes and their separation by spacers can be observed directly by electron microscopy in gently lysed and dispersed nuclei.
The hundreds of genes for rRNA form the nucleolar organiser, the core of the nucleolus. In the nucleolus rRNA transcripts are processed into three ribosomal RNAs and combined with the fourth rRNA (5S) and with ribosomal proteins, which are synthesized in the cytoplasm and imported into the nucleus, to form the large and small ribosomal subunits. These are then exported to the cytoplasm.
Thus, the nucleolus is the structural manifestation of rRNA transcription, rRNA processing, and assembly of ribosomal subunits. Genes encoding 5S rRNA are arranged in multiple tandem clusters, usually in several different chromosomes and do not form part of the nucleolus. In Xenopus there are 24,000 copies of 5S genes that are separated by un-transcribed spacers.
The genes are 120 bp long and are separated by spacers of 600 bp. Genes encoding tRNA molecules are typically repeated about 200 times and are separated by spacers that are, on the average, 10 times longer than a tRNA gene itself. In growing oocytes of amphibians and some insects even the many copies of genes encoding 18S, 28S, and 5.8S rRNAs are insufficient to meet the need for ribosome production.
In these cells the genes for these rRNAs are extensively amplified further. Histones are proteins needed in the packing of long DNA molecules in the nucleus. There are five types of histones and 20 to several hundred copies of each gene are present.
In most eukaryotes histone genes are arranged in groups, with one copy of each of the five genes per group. In sea urchin cells there are about 1000 such groups repeated in tandem. The five genes within a group are transcribed individually and separated by spacers. The groups in turn are separated from one another by spacers.
The spacers that occur between copies of repeated genes are usually similar or identical in sequence for a particular gene in a particular species. Thus, all the spacers between 5S rRNA genes are the same in Xenopus laevis.
In the closely related species, X. mulleri, the 5S genes are identical to those in X. laevis but the spacers are three times longer and have a different sequence from the spacers in X. laevis. In general, spacers change in length and sequence far more rapidly during evolution than the genes they separate. Whatever the function of spacers, that function is compatible with large changes in spacer composition.
The significance of spacers is unknown. However, it is presumed that spacers between repeated genes represent the same phenomenon and have the same significance as spacers between single copy genes.
Packing of DNA in Chromosomes:
The length of the DNA molecules in eukaryotic chromosomes exceeds by many thousands the diameter of the nuclei in which they are contained. Extreme examples are the DNA molecules of some salamanders. Twelve DNA molecules, each several meters long, are contained in a spherical nucleus less than 20 Am in diameter in Amphiuma.
In humans a total of 200 cm of DNA in the 46 chromosomes of a diploid cell are contained in a nucleus with a diameter of 6 or 7 μm. By coiling and packing, the extremely long DNA molecules can fit into the small nuclear volume. Certain nuclear proteins, bring about the coiling and packing process.
Nucleosomes and Chromatin:
The DNA of eukaryotes is complexed with proteins that, in effect, package the DNA in stretches and hold it in specific configurations within the chromatin. The arrangement of proteins and DNA in chromatin is partially understood. Also, it is clear that the particular configuration of DNA in chromatin is very important in the regulation of transcription.
In general DNA packaged tightly with protein in chromatin is not transcribed; loosening of the chromatin is necessary, but alone is insufficient to bring about transcription. For transcription to occur other proteins must interact with the DNA sequence upstream from the coding region of a protein-encoding gene.
The packing of DNA in chromatin occurs in several successive orders, each of which accomplishes a further compaction of the DNA. The primary level of packing has been thoroughly elucidated by biochemical and physical analyses and electron microscope observations on chromatin isolated from interphase nuclei. It has been known for a long time that DNA is complexed with a family of five histone proteins.
Histones contain large numbers of the basic amino acids lysine and arginine, with R groups that terminate in amino groups (—NH2). In the pH range in the cell the amino groups acquire a proton (H+) and are therefore charged positively (—NH3+).
The large number of positive charges of arginine and lysine residues in histones form, ionic bonds with the negative charges of the phosphates in the DNA strands. In the last decade the molecular details of how histones are complexed with DNA and form the first level of chromatin organisation have been worked out.
Brief treatment with deoxyribonuclease (DNase) breaks chromatin into small particles, each composed of histones complexed with a fragment of DNA. After the histones are removed, the DNA consists of fragments of about 200 by and multiples of 200 by (i.e., 400, 600, etc.). If the enzymatic digestion is longer, all the DNA is in units of 200 bp, and the multiple-size fragments are not found. These and other experiments show that the long, continuous DNA molecule in a chromosome is complexed with histones in 200-bp stretches.
Each stretch is complexed with histones so as to form a particle. In corollary observations by electron microscopy, chromatin that has loosened and spread out is found to consist of long strings of particles. Brief treatment with DNase breaks the string into units containing one to several particles, and longer digestion reduces the string entirely to individual particles. Each particle can be dissociated into a 200-bp fragment of DNA and a particle consisting of eight histone molecules.
The particles of chromatin, or nucleosomes, are joined in long strings because the DNA molecule is continuous from one nucleosome to the next. The region of DNA linking nucleosomes is more accessible to DNase than DNA within nucleosomes. Hence, with properly gauged digestion chromatin is converted into single, physically separate nucleosomes.
Each of these nucleosomes is made up of a 200-bp fragment of DNA and nine histone molecules-one molecule of histone H1, and two molecules each of the other four histones H2A, H2B, H3, and H4. If the nucleosome preparation is digested further with DNase, the DNA fragment is gradually reduced in size until it reaches 146 bp and histone H 1 is released. Further action of DNase is blocked because of a tight association between the remaining eight histone molecules and DNA.
The structure that remains is called the core particle. The eight histone molecules (two each of H2A, H2B, H3, and H4) in a core particle are joined with each other in a specific pattern that forms a disc. The DNA molecule makes almost two turns around the disc and then continues to the next core particle. The DNA extending from one core particle to the next is called linker DNA.
While the amount of DNA wrapped around the histone aggregate is constant (146 bp), the amount of DNA in the linker is variable from one kind of cell to another or from species to species. On the average, linkers are a little over 50-bp long, which, added to the DNA around the core particle (146 bp), accounts for the 200-bp fragments observed after brief DNase digestion of chromatin.
An H1 histone molecule is attached to each stretch of linker DNA; hence it is released when the linker DNA is digested with DNase. H1 histone plays a role in holding adjacent nucleosomes in tight association and in folding or twisting of the string of nucleosomes into the next order of DNA packing.
The packing of DNA into nucleosomes condenses the DNA about seven-fold, forming on the average for a human DNA molecule (four cm long) a thread of 600,000 nucleosomes that is about ten nm (1 nm = 10-9 meter) in diameter and 6000 Am long. Relative to the diameter of the nucleus (6 to 7 nm), the string of nucleosomes is still enormously long.
In the next order of DNA packing, the ten-nm nucleosome thread is folded into a fiber about 30 nm in diameter. According to the most widely accepted model the nucleosome thread is arranged as a hollow helix or solenoid, generating a thicker structure.
The solenoid is believed to account for the thread of 30 nm in diameter seen by electron microscopy of chromosomes. The thicker filament formed in the solenoidal model is about seven times shorter than the extended nucleosome filament or a little over 800 t4m for an average human chromosome.
Observations by electron microscopy suggest that at the next higher level of packaging the solenoidal filaments are arranged as loops, containing 5 to 10 × 104 by of DNA in an animal cell, extending out from a central axis composed of protein and resembling the arrangement seen in lampbrush chromosomes.
The three orders of packing of DNA (nucleosomes, solenoids, and central axis with loops) accomplished by histones and other proteins is thought to produce the chromatin observed in the interphase nucleus. This chromatin contains the DNA that is transcribed and forms euchromatin.
A portion of the chromatin undergoes additional packing to yield the more tightly condensed form, heterochromatin. The molecular arrangements in this mode of packing are not known. The significance of euchromatin vs. heterochromatin is not completely understood, but DNA present in heterochromatin is generally not transcribed.
Organisation of Chromatin:
The contents of the nucleus appear to form a gel with no movement of the euchromatin, heterochromatin, or nucleolus. There is, for example, no Brownian movement (vibratory movements caused by molecular collisions) of intranuclear structures in living cells viewed by light microscopy.
By comparison, the cytoplasm is largely a solution in which organelles like mitochondria and lysosomes visible in the light microscope are in constant Brownian movement. The immobilised state of interphase chromosomes also contrasts sharply with the constant Brownian movement activity and shuffling movements of chromosomes in mitosis.
Those parts of interphase chromosomes that are condensed into heterochromatin are often closely associated with the inside of the nuclear envelope. A striking example is the inactivated X chromosome in a female mammalian cell. The entire chromosome is packed as a discrete mass of heterochromatin, the Barr body that is also positioned at the nuclear envelope.
The inside of the nuclear envelope is lined with a thin layer of protein, the nuclear lamina. The lamina may serve to hold and therefore probably mediate the adhesion of chromatin to the nuclear envelope. The two membranes of the nuclear envelope are continuous with one another as they form the walls of the many nuclear pores that are present in the nuclear envelope.
The inner and outer openings of a pore and the interior of the pore contain eight granules, arranged radially and surrounding a central granule that is sometimes present in the opening. The granules, together with the pore, are called the nuclear pore complex. The entire complex is about 100 nm in diameter. The particles of the pore complex together with some ill-defined fibrous material appear to occlude most of the pore opening.
Nuclear pores are nevertheless believed to be channels by which materials move between nucleus and cytoplasm. When molecules with molecular weights of less than 50,000 are injected into the cytoplasm, they quickly equilibrate between the nucleus and cytoplasm. Larger molecules enter the nucleus slowly. From such experiments it has been estimated that the effective diameter of the opening of the pore is about nine nm.
Molecular traffic through nuclear pores is intense. For example, during DNA replication hundreds of thousands of histone molecules synthesised in the cytoplasm enter the nucleus. Ribosome production in the nucleolus requires the continuous importation of great numbers of the approximately 80 different kinds of ribosomal proteins.
Experiments suggest that histones and other proteins may be actively transported through the pores. In addition, large numbers of mRNA- protein complexes and new ribosomal subunits flow out of the nucleus every minute. These substances are too large to be accommodated by the nine-nm opening of a pore, and some means of increasing the effective opening to facilitate their passage must occur.
The particle sometimes observed in the center of a nuclear pore may represent material caught in transit between nucleus and cytoplasm. During mitosis in most kinds of cells the nuclear envelope breaks into small fragments that disperse in the cytoplasm. These come together again to reform nuclear envelopes around daughter nuclei in telophase.
The specific re-aggregation of these fragments shows that the membranes of the nuclear envelope must have properties that distinguish them from membranes of the endoplasmic reticulum. The re-formation of the nuclear envelope is closely coupled with the re-formation of the nuclear lamina on the nuclear surface of the inner membrane.
Structure of Condensed Chromosomes:
As a cell enters prophase of mitosis or meiosis the interphase chromatin becomes tightly condensed by further packing, and individual chromosomes become recognisable. An early stage of such packing is represented by the lampbrush chromosome, in which the central axis has probably shortened and thickened compared to the interphase state.
The central axis is, in turn, folded or coiled to produce the fully condensed chromatids of a chromosome. The condensation of chromosomes is achieved by proteins. The synthesis of a principal protein involved in condensation begins just around the start of prophase, but details about the events of condensation are still lacking.
Removal of histones and DNA from metaphase chromosomes leaves a residual structure that is still recognisable as a chromosome. The proteins that are left are believed to form the scaffold onto which the DNA histone components are organised and packed into the condensed mitotic or meiotic form.
The chromosome scaffold and the nuclear matrix of the interphase nucleus may well be forms of the same structure, one designed to produce the compact mitotic chromosomes, the other responsible for the organisation of the transcriptionally active interphase chromosome.
Banding in Mitotic Chromosomes:
Mitotic chromosomes are not uniform in structure throughout their lengths; a considerable amount of substructure is revealed by a variety of staining techniques. The substructure takes the form of several kinds of transverse bands. Harsh treatments that extract DNA differentially from regions of a chromosome leave certain regions intact. In subsequent staining, these regions, known as C-bands, show up more intensely.
C-bands in general correspond to DNA that is packed as heterochromatin in the interphase nucleus. In metaphase chromosomes, this DNA appears to be more tightly packed than DNA that occurs as euchromatin during interphase. C-banding occurs predominantly in the centromeric region of mammalian chromosomes, where heterochromatin containing highly repetitive DNA is known to be located.
Three other staining techniques produce R-, Q-, and G-bands. The different patterns of C-, R-, Q-, and G-banding are constant in a species and allow each chromosome to be identified unequivocally Figure 8.27. shows the banding map for the chromosomes of the human genome.
Banding in mitotic chromosomes is an entirely different phenomenon from the banding seen in polytene chromosomes. The latter are interphase chromosomes and their banding should not be confused with the banding in mitotic chromosomes. Banding patterns in mitotic chromosomes are useful because they allow segments of chromosomes that have been exchanged between chromosomes (translocations) to be precisely identified.
Translocations can be induced by radiation and sometimes they occur without known cause. They occur frequently in cancer cells, and often they have important consequences for the cell.
Replication of Chromosomes:
In most eukaryotes the DNA molecules of chromosomes are enormously long, raising the question of how replication is organised. Prokaryotic chromosomes, which are shorter and circular, replicate beginning at a single origin; that is, they are single replication units. In contrast, the linear DNA molecule of a eukaryotic chromosome contains many points of initiation of replication; that is, it contains many replication units.
Replication Units:
The multiple replication units of eukaryotic chromosomes are readily seen by autoradiography or by electron microscopy. The autoradiographic experiment is simple. Cultured cells are incubated with 3H- thymidine for a few minutes during DNA replication. Cells are lysed with a drop of a solution of an ionic detergent on a microscope slide. The detergent lyses the nuclei and removes essentially all proteins from the DNA. The drop of lysed material is spread evenly over the entire microscope slide.
This draws the DNA molecules out in long, linear stretches. Subsequent autoradiography shows that DNA replication was occurring at many places along individual molecules during the short incubation with 3H-thymidine. Each stretch of radioactivity represents a replication unit. Replication can also be seen directly by electron microscopic observation of isolated DNA molecules. Each segment of the molecule that is doubled represents a separate replication unit.
In prokaryotic chromosomes origins of replication are defined by specific deoxynucleotide sequences. The strongest evidence that origins are similarly defined in eukaryotes comes from the study of yeast chromosomes. ARSs that function as apparent specific origins of replication in yeast plasmids have been identified in the DNA molecules from a variety of eukaryotes. Conclusive evidence that these sequences function as origins of replication in the chromosomes of these eukaryotes is still lacking.
Bidirectional Replication:
Replication is initiated at the single point within each replication unit and proceeds bidirectionally from that point much as in prokaryotes. Such points are labeled as origins, but whether they represent specific sequences has not been unequivocally proven. Bidirectional replication was discovered by the following autoradiographic experiment- fluorodeoxyuridine, which inhibits DNA replication, was used to block entry into the S period. A culture of cells was thus synchronised. Cells in the G2, M, and G1 periods of the cell cycle are unaffected by the drug, proceed to the G 1 /S transition point, and stop.
Cells already in S are killed by fluorodeoxyuridine. After enough time was allowed for cells to be synchronised at the G 1/S border, the inhibitor was washed away and fresh nutrient medium containing 3H- thymidine was added. As a result, replication was initiated, with incorporation of 3H-thymidine. After a few minutes of replication, nonradioactive thymidine was added to the medium. The thymidine was taken up by cells and phosphorylated to TTP, which became part of the pool of TTP used for DNA replication.
Over several minutes nonradioactive TTP replaced 3H-TTP in the pool of TTP in the cell. As a consequence the amount of radioactivity incorporated into DNA progressively diminished. Some minutes after addition of nonradioactive thymidine, cells were lysed; autoradiography of DNA molecules showed the expected tandem arrays of replication units.
However, each autoradiographic image was darkest in its central region and faded to imperceptibility at its two extremities. The fading images reflect the travel of replication forks during the decline of 3H- thymidine triphosphate in the cell and show that replication has preceded bidirectionally from a single point (origin) in each replication unit.
Termination between Replication Units:
Each replication fork eventually meets a replication fork traveling toward it in the next adjacent replication unit. Meeting of forks completes replication of that particular segment of the chromosome, and the replication machinery dissociates from the DNA. The terminus between adjacent replication units is defined by the meeting place of replication forks rather than by a specific sequence in the DNA.
The meeting of forks and completion of replication between replication units leaves the two daughter molecules linked by one interloop between them for each replication unit. These interloops would be difficult to eliminate by untwisting of the two duplexes at the end of replication. An average of more than 1300 interloops is generated in a human chromosome.
Loops are removed by the action of topoisomerase, which breaks the chains in the daughter double helices, allows the daughters to separate and then reseals the breaks. Mutant yeast cells lacking the proper topoisomerase cannot remove interloopings between the two daughter molecules. As a result, the two chromatids in a chromosome cannot separate from each other at the ensuing mitosis.
Size and Number of Replication Units:
An estimate of the size of replication units is obtained by measuring the distance between the centers of successive origins either in autoradiographic or electron microscope displays of DNA. Replication units in mammalian cells range from 15 to 100 μm (45,000 to 300,000 bp) with an average of 30 p.m (90,000 bp). The four cm of DNA in an average mammalian chromosome therefore contain over 1300 replication units. A haploid set of mammalian chromosomes contains about 33,000 replication units.
Rate of Replication:
The rate of travel of replication forks has been estimated by several methods, among them autoradiography. The simplest procedure is to synchronise cells by blocking them at the G 1/S border, then releasing the block and simultaneously adding 3H- thymidine. DNA isolated at intervals thereafter produces progressively longer autoradiographic images, which shows that in mammalian cells replication forks travel about one µm per minute (3000 bases per minute). Since the average size of a replication unit is 30 μm and each unit has two forks, the time to replicate an average-sized unit is 15 minutes.
4. Term Paper on the DNA Molecules in Mitochondria:
The mitochondria of all eukaryotes contain DNA molecules (mtDNA). These encode a few of the macro-molecules that make up mitochondria, although all but a few of the 100 or more different kinds of polypeptides in mitochondria are encoded by nuclear genes. The polypeptide products of these nuclear genes are synthesized in the cytoplasm and subsequently imported into mitochondria.
Among protozoa, yeasts, and other unicellular eukaryotes mitochondrial DNA molecules range in size from 15,000 to 75,000 bp. In a few species the molecules are linear, but in most they are circular. There are multiple identical copies of mtDNA molecules in each mitochondrion, and all molecules in all the mitochondria have the same coding functions.
Among unicellular eukaryotes mtDNA of the budding yeast has been studied intensively. It is a circular molecule containing 75,000 bp, and there are ten to 50 DNA molecules altogether in all the mitochondria of a single cell. These mtDNA molecules represent 5 to 25 percent of the total cellular DNA but only 0.5 percent of the total sequence complexity of the total DNA.
These are unusually high percentages, and are due to the unusually large size of yeast mtDNA (75,000 bp) and the small size of the yeast nuclear genome (1.5 × 107 bp). In all other eukaryotes studied so far, mtDNA represents a far smaller percentage. In mammalian cells mitochondrial DNA makes up less than one percent of the total cellular DNA and about 0.0005 percent of sequence complexity of the total DNA in the cell.
Molecules of mtDNA in invertebrate and vertebrate animals are closed circles as in yeast but are much smaller, ranging in size from 13,000 to 18,000 bp. In mammalian mitochondria, mtDNA molecules contain about 16,000 bp.
The entire sequence of 16,569 by in human DNA has been determined. There are five to ten molecules per mitochondrion and 100 to several thousand mitochondria per cell, depending on the size and type of cell. Fibroblasts have about 100, liver cells about 1000, and heart muscle cells many thousands.
In plants, mtDNA molecules are much larger and more variable in size than those in animals. Even within a single organism the lengths of mtDNA molecules are variable, and some molecules are circular and some are linear. These variations are apparently not the result of DNA breakage during preparation and are not understood.
Much less is known about the coding functions of mtDNA in plants compared with yeast and animal mtDNA, but evidence so far suggests that plant mtDNA contains about the same number of different kinds of genes as other eukaryotes.
ADVERTISEMENTS:
Mitochondrial DNA molecules are not complexed with nuclear- type histones to form nucleosomes. Histone-like molecules are present instead, and die mtDNA-protein complexes form nucleoid bodies in the matrix compartment.
Mitochondria reproduce by fission, and the multiple mtDNA molecules, which are located in the matrix of mitochondria, are distributed randomly between daughter mitochondria. In some kinds of cells, for example, chick cells in culture, and probably in cells in general, mitochondria fuse and separate often, so that the population of mtDNA molecules is dynamically shared among the mitochondria within a single cell.
A yeast mtDNA molecule is five times longer than a mammalian mtDNA molecule but both encode almost an identical set of genes. Most of yeast mtDNA has no known coding or other function. MtDNA molecules in both organisms encode three of the subunit polypeptides of cytochrome c and seven or eight other polypeptides.
Both encode the two major rRNA molecules for large and small subunits of ribosomes. These rRNAs are smaller than those encoded by nuclear genes, and mitochondrial ribosomes are smaller than cytoplasmic ribosomes. Mammalian mtDNA encodes 22 kinds of tRNA molecules and yeast encodes about 25.
Thus, mtDNA encodes some of the molecules needed to make up the translation machinery in the mitochondrial matrix. However, most elements, like most ribosomal protein and aminoacyl-tRNA synthetases, are encoded by nuclear genes, synthesized in the cytoplasm, and imported into the mitochondrion.
The mitochondrial translation machinery is limited in use to the several kinds of mRNA molecules transcribed from mtDNA. No mRNA molecules are imported from the cytoplasm. Like bacterial mRNA molecules, mitochondrial mRNAs are not capped at their 5′ ends. They have short, 3′-poly (A) tails.
Transcription of human mtDNA is unusual. One long transcript is produced from each strand of the DNA, in contrast to transcription from only one strand in nuclear DNA. The two transcripts are full- length copies of the two DNA strands. Both transcripts are processed, each yielding some of the rRNA, tRNA, and mRNA products. The coding segments for some genes are in one strand of the DNA and coding segments for others are in the opposite DNA strand.
Remarkably, the genetic code of mtDNA varies slightly from the nearly universal code present in the nucleus of eukaryotes, the nucleoid of prokaryotes, and viruses. In fungi, which includes yeast, and in animal cells, the codon UGA functions as a codon for tryptophan rather than as a stop codon for translation. In the mtDNA of some organisms AUA codes for methionine rather than for isoleucine. AGA and AGG code for stop rather than arginine.
During the cell cycle the total number of mtDNA molecules doubles, keeping pace with a doubling in mitochondrial number. Replication is catalysed by a mitochondrial-specific polymerase called DNA polymerase y, which is encoded by a nuclear gene. The mtDNA molecules replicate asynchronously with each other. Some molecules may replicate twice during a given cell cycle and others not at all. The time of replication is spread over the entire interphase and bears no relationship to the nuclear S period.
At least some segments of mtDNA are represented by nearly identical sequences at scattered locations in the nuclear genome. For example, the 16S rRNA sequence of human mtDNA is also present in nuclear DNA. The significance of mitochondrial sequences in the nucleus is not known.
5. Term Paper on the DNA Molecules in Chloroplasts:
Chloroplast DNA molecules (chl DNA) range in size from 33,000 by to more than 2 × 106 by. In higher plants chl DNA molecules consist of about 1.5 × 105 bp. The number of molecules per chloroplast ranges from 20 to more than 100, depending on the species, and these are distributed in multiple nucleoid-like bodies in the stromal compartment of the chloroplast.
More than 80 polypeptides are encoded by chl DNA, as well as rRNAs for large and small ribosomal subunits and tRNAs. Many other chloroplast proteins are encoded by nuclear genes, are synthesized in the cytoplasm, and are imported into the chloroplast. For example, the small subunit polypeptide of the enzyme ribulose-1, 5-bisphosphate carboxylase is encoded by a nuclear gene; the large subunit polypeptide is encoded by chl DNA.
This enzyme occupies a pivotal position in the conversion of CO2 to carbohydrate during photosynthesis. Messenger RNA molecules transcribed from chl DNA are translated by the chloroplast translation machinery in the stromal compartment of chloroplasts.
Like mtDNA molecules chl DNA molecules double in number during the cell cycle and are distributed randomly between the daughters during chloroplast fission. Histones are not associated with chl DNA.