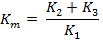ADVERTISEMENTS:
The following points highlight the top four types of chromosomal aberrations. The types are: 1. Deficiency or Deletion 2. Duplication 3. Translocation 4. Inversion.
Chromosomal Aberration: Type # 1. Deficiency or Deletion:
Chromosomes contain a number of genes on them. The genes are arranged in linear fashion. The number and also the positions of genes on a chromosome are fixed. When a chromosome is broken into several pieces, the healing (reunion) of the segments takes place and it is possible that the two ends of the fragments unite together leaving one or more acentric parts free (Fig. 22.3).
The acentric pieces of chromosomes as mentioned earlier disappear. The healed segments will form a chromosome that will be deficient for some of the genes, particularly those found on the lost part. Depending upon the length of chromosome segment lost in this way, the loss involves one gene or a block of genes.
ADVERTISEMENTS:
The loss of a section of genetic material and genetic information from a chromosome or linkage structure is termed deficiency or deletion. Deletions may be produced in several ways, such as by loss of terminal acentric fragments or interstitial segment of chromosomes Fig. 22.3).
Although in practice deletions frequently refer to losses of terminal as well as intercalary segments of chromosome, at molecular level the deletions, of course, can be so small that many mutant loci are in reality deficient for one or more nucleotides in the DNA molecule. If the deficiency is large, the chances are there that the cell may die.
However, if the deficiency is small, the cells may persist. Should such a cell be fertilized egg, the net effect will be production of a dominant lethal. Homozygous deletions are usually lethal but heterozygous deletions appear as normal mutations.
ADVERTISEMENTS:
Deficiency can be detected by its two characteristics, namely, genetic effects and cytological effects.
Genetic Effects of Deficiency:
These are primarily due to the loss of genetic information and secondarily due to qualitative changes in the genotype as well as the change of genie balance. Deficiencies have been useful in determining the exact locations of genes on the chromosomes.
One of the genetic effects of deficiency is pseudo-dominance. Pseudodominance occurs when the chromosome that carries the deletion of dominant allele pairs with a normal homologous chromosome carrying the recessive allele.
In absence of dominant alleles the recessive alleles would be expressed in phenotype as if they were dominant. This is called pseudo-dominance. If some of the corresponding genes have tendency of lethal effect in double dose, they will be lethal in single dose (recessive lethality). The chromosomes with deletion never return to a normal state.
Cytological Effects:
The existence of relatively large deletions in the chromosome complements of eukaryotes may be proved cytologically
(i) By the occurrence of centric and acentric chromosome fragments in mitosis, and
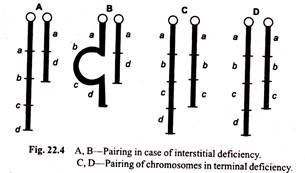
(ii) By the absence of regional pairing during first meiotic prophase. If the cell is heterozygous for deletion, i.e. it has a normal chromosome and a deficient homologue, then during synapsis, the chromosomes pair precisely gene by gene all along the homologous region and in deficient region, however, the normal chromosome will not pair.
Therefore, a buckling (loop) will develop in the normal homologue at the point of intercalary deficiency (Fig. 22.4). It is, therefore, known as buckling effect. Such a configuration is called deficiency loop or compensation loop and can be observed under microscope.
The pseudominance and the buckling in synapased chromosomes can thus be used to determine the exact location of genes in the deficient region. The genes which are known to be missing in the deficiency must be located in the buckle or loop region of homologue.
ADVERTISEMENTS:
An example of deficiency is known in X-chromosomes of Drosophila in which few bands are missing from the tip of X-chromosome. Such a deficiency characterised by heteromorphic bivalent during prophase of first meiosis and has well marked phenotypic effect in Drosophila.
Notches in the wing margins of female fly are actually due to this sort of deletion in X-chromosomes. It is lethal in males. Many of the minute (bristles) series of mutants in this fruit fly are also due to deficiency.
In the lower organisms, for example, Chlamydomonas, yeasts and some fungi, deficiencies are totally lethal, i.e., they result in the death of the individuals.
ADVERTISEMENTS:
Chromosomal Aberration: Type # 2. Duplications:
A structural change resulting in the doubling of genes in a section of the chromosome of prokaryotes and eukaryotes is referred to as duplication. In other words, the inclusion of extra part or duplicated gene sequence of a chromosome beyond the normal complement is called duplication.
If a segment of one chromosome is incorporated in another homologous chromosome, it is called intrachromosomal duplication (Fig. 22.5), but if the duplicated chromosome segment is either incorporated into a non-homologous chromosome or occurs as a fragment in the chromosomal set it is called interchromosomal duplication (Fig. 22.5).
The duplicated segments contained within a single chromosome could exist in one of the following configurations depending upon the position and sequence of the duplicated genes.
(i) Direct tandem duplication in which the duplicated gene sequence lies just next to normal corresponding section and the sequence of genes with respect to centromere is the same in duplicated segment as in the normal section of the chromosome.
It will be clear from the following gene sequences in the normal and duplicated chromosomes:
(ii) Reverse tandem duplication in which the duplicated section with reverse gene sequence lies adjacent to normal sequence as shown below:
(iii) Displaced direct duplication in which the duplicated section is not adjacent or contiguous with the normal section (i.e., separated by other segment).
The duplicated and normal gene sequences may be in the same arm (intra arm) or in two different arms of the chromosome (inter arm duplication):
(iv) Displaced Reverse duplication in which the duplicated section with reverse gene sequence is separated from normal segment by other segment as shown in the following chromosomes:
(v) Transposed duplication in which the duplicated gene sequence is attached to another position owing to inter-chromosomal duplication.
ADVERTISEMENTS:
The size of duplicated segment may vary considerably. The smallest duplications that can be investigated cytologically are those of single band of polytene chromosomes. However, the whole arm of the chromosomes may be duplicated, thus giving rise to isochromosome. Where all linkage groups or chromosomes in the haploid chromosome set are doubled, this is referred to as genome mutation.
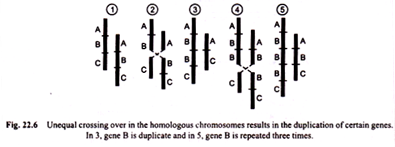
Duplication may arise in several ways. One of the most common methods is unequal crossing over, a process which produces one chromosome with duplication and another with deficiency. This is primary structural change of chromosome.
The schematic representation of gene duplication in chromosome by unequal crossing over is shown in Fig. 22.6.
Duplications may also occur due to crossing over in inverted or translocated segments. (Secondary structural mutation of chromosome)
ADVERTISEMENTS:
Duplications in general, are much more viable than deficiency. A heterozygous duplication has an appearance similar to deletion. Duplications sometimes appear as dominant mutations. By duplication an allele may be duplicated, triplicated or multiplicated, hence duplications may be utilized a studying the effect of various quantitative doses of members of an allelic set.
Well known example of duplication which had a significant impact on genie theory is Bar-eye mutation in Drosophila. With the discovery of chromosomal nature of this case, it was found that extra pieces of chromosome were associated with a normal X-chromosome duplicating and triplication section of it (Fig. 22.7).
The Bar-eye, a sex-linked incompletely dominant mutation responsible for the development of rod-shaped eye with reduced eye facets, appeared spontaneously in a wild type stock of Drosophila melanogaster with round eyes. Homozygous stock of Bar-eyed mutants produced flies with normal eye and flies with even more reduced eye (Double Bar) in approximately equal frequency.
These observations suggested that Bar locus was very unstable but the appearance of wild type and double bar flies in equal numbers could not easily be explained. Moreover, the two mutant phenotypes appeared in the progeny when Bar-females were crossed with normal males, and not in the progeny of Bar-males crossed with normal females.
Drosophila is unusual in the sense that meiotic crossing over occurs only in females and not in the males. This observation suggests that mutation was not responsible for the wild type and double Bar progeny and that Bar might somehow be related to meiotic crossing over.
In 1925, Sturtevant using females homozygous for Bar (B) but heterozygous for forked bristles gene f) and for fused wing veins (gene fu) demonstrated that the normal and double Bar flies were products of crossing over within the Bar locus. Sturtevant found that some normal and some double Bar flies were recombinants, each with one or the other flanking marker (‘f’ or ‘fu’).
Sturtevant hypothesis of unequal crossing over was supported by Bridges in 1936. Bridges studied the chromosomes of wild type Bar eyed and Double Bar eyed flies and noticed the presence of single 16A segment in normal eyed, two 16A segments in tandem in Bar eyed and three continuous 16A segments in double Bar eyed (Fig. 22.7). By unequal crossing over in homozygous Bar females one 16Å.
Segment is transferred to a chromatid with duplication and consequently a chromosome with only a single 16Å segment (normal type) and another with three 16Å segments (Double Bar) are recovered. Duplication and reduplications of Bar region can be easily seen in the salivary gland chromosomes (Fig. 22.7).
The chromosomes of Drosophila salivary gland are large and easily distinguishable. They replicate several times but do not divide. So, each chromosome appears like a ‘rope’ of about 1,000 tightly coiled chromatids—a polytene chromosome. The genes for Bar eye and several other characters have been, in fact, cytologically pinpointed to specific bands or chromomeres.
In com and peas, a number of duplicating factors are known. Duplications like deletions may be so small as to be molecular. Since they change the genie balance, they may produce abnormalities m body characters. Duplications are considered to play a role in origin of new genes through functional diversification of duplicated members.
Like deficiencies, duplications have diagnostic cytological and genetically effects.
Cytological Effects:
The existence of relatively small duplications in the chromosome complements of eukaryotes may be proved by the appearance of regional distortions of duplicated chromosome during pairing in first meiotic prophase or somatic pairing in specialized tissue like salivary glands of Diptera.
Crossing over in reverse tandem duplication may result into a dicentric chromosome. This can be frequently observed in maize. As the two centromeres move to opposite poles, a chromosome bridge is formed which later on breaks at any point along the bridge.
The broken ends are sticky and the replication of the broken pieces may result in two sister chromatids which may be joined together due to their sticky ends. Thus, whole chromosome arm may be duplicated giving rise to isochromosomes. In sporophytic tissues of plants, the isochromosomes are uncommon.
Genetic Effects:
The genetic effects of duplications depend on the genetic information the duplicated segments contain and the change in genie balance effected by them. In homo and heterozygous state they may cause an increase or decrease in the viability of their carriers and in extreme cases may act as lethals.
Since the duplications supply the additional genetic material and change the genie balance, they play important role in evolution at individual and population levels.
Under evolutionary conditions, small duplications may provide a basis for the mutational differentiation of genetic material and the different copies of the same gene may change in different directions without disturbing the normal functions of an organism.
The cases in which different gene pairs affect the same character (as for example, multiple factors, complementary factors) possibly arose initially after duplication of single genes. The repetition of DNA sequences frequently seen in highly evolved organisms is a direct indication to this.
Chromosomal Aberration: Type # 3. Translocation:
Chromosomes may break into two or more fragments, each with a raw end. These segments may reunite and during reunion either the pieces of the same chromosome or the pieces of the non-homologous chromosomes may be fused. Ionising radiations such as X-rays and gamma-rays are frequently used to break chromosomes for producing structural changes.
In this process, a part of the chromosome is transferred to another non-homologous chromosome within the chromosome complement.
In other words, a translocation is a chromosomal rearrangement which involves:
(i) The unidirectional transfer of a chromosome segment and its gene sequence to a different chromosome within the chromosome complement, and
(ii) The exchange of segments between non-homologous chromosomes. In this way, translocation only results into a change in the sequence and position of genes, their quantity being unaffected.
Types of Translocation:
1. Simple Non-Reciprocal Translocations or Transpositions:
In this process, a piece of one chromosome is transferred to a non-homologous chromosome.
It can be of two types:
(i)Simple translocation or Terminal transposition,
(ii) Shift translocation or Interstitial transposition.
(i) Simple Translocation:
When a chromosome breaks into two parts due to external or internal stress, one of the two segments of the broken chromosome may become attached to the natural end of the nearest chromosome which may not be its homologue. This is simple translocation.
Suppose, there are two non-homologous chromosomes A B C D E F G and TUVWXYZ. If F G part of the first chromosome is transferred to second chromosome, a new chromosome TUVWXYZFG would result as shown in Fig. 22.8. In this non-standard or translocation chromosome, F G is said to be in simple translocation state.
In the normal course, the terminal transpositions are not common because the raw ends or the telomeres of unbroken chromosomes are not sticky.
(ii) Intrachromosomal, Shift Translocation or Interstitial Transposition:
In this, an interior or interstitial segment of a chromosome that is induced by two breaks is incorporated interstitially into another non-homologous broken chromosome, the latter being induced by a single break. It is also called intercalation or insertion.
If the sequence of genes in the translocated segment is the same as that in the original segment with respect to centromere, it is referred to as encentric translocation. But if the sequence of gene loci in translocated chromosome segment is reversed it is called dyscentric translocation.
2. Reciprocal Translocation:
When a break occurs at a point where two non-homologous chromosomes touch each other, the broken end of one chromosome may become united with the broken end of second chromosome and that of second chromosome becomes attached to that of first, this is reciprocal translocation.
Suppose, there are two chromosomes A B C D E F and G H I J K L. These two after reciprocal translocation may produce chromosomes A B C J K L and G H I D E F as shown in Fig. 22.9. The reciprocal translocation is the most common type of translocation.
If there occurs two breaks in each of the two non-homologous chromosomes, the reciprocal translocation of intercalary segments may be obtained, but it is very rare.
The reciprocal translocation is like crossing over except that it involves exchange between the segments of two non-homologous chromosomes. It is sometimes called “illegitimate crossing over”.
The origin of translocations is interpreted either according to the breakage-reunion or the exchange model. The unit of translocation may be the chromosome (chromosome translocation) the single chromatid (chromatid translocation) or a segment of chromatid. The sites of translocations are called translocation points. The reciprocal translocation may be either asymmetrical (aneucentric) or symmetrical (eucentric).
The asymmetrical translocation gives rise to one dicentric and one acentric chromosomes (Fig. 22.10) and it may lead to the formation of chromosome bridge if the two centromeres of a dicentric translocation product are distributed to opposite poles of spindle during anaphase. In symmetrical translocation, however, the products are monocentric.
Sometime, whole or nearly whole arms of the chromosomes may be transposed or interchanged. This is termed whole-arm translocation (Muller, 1940).
There are three special cases of whole arm translocation:
(a) Centric fusion,
(b) Dissociation, and
(c) Tandem fusions.
(a) Centric fusion:
Translocation occurring between two sub-telocentric chromosomes, each with sub-terminal centromere and single long arm, may exchange parts in such a way that major part of long arm of one is translocated to the short arm of the other producing one V-shaped long metacentric chromosome (i.e., with median centromere) and one small chromosome with two minute arms.
Long V-shaped chromosome has all the vital genetic material and the small chromosome is little more than a centromere plus some chromatic material.
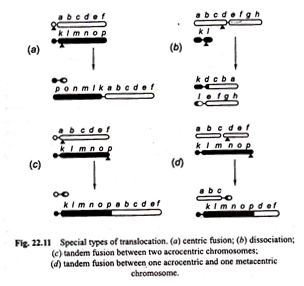
This type of translocation is called “centric fusion” by White (1954) [Fig. 22.11 (a)]. Such a translocation may be quite viable and in a heterozygote, one long V-shaped chromosome may synapse normally with two normal unfused one armed homologous chromosomes.
The small chromosome is more or less functionless and appears like a supernumerary one which may be lost from any individual but maintained in the population. These supernumerary chromosomes are spare kinetochores.
The reverse translocation in them may lead to increase in basic chromosome number. It seems a very likely mechanism for the alteration in the chromosome number, although there is little direct evidence for it.
In general, the translocation appears to be major method of genome alteration. Thus it leads to reshuffling of genie loci as well as alterations of basic chromosome morphology. Centric fusions taking place between a sex chromosome and an autosome may represent the origin of “multiple sex chromosome” system.
(b) Dissociation:
Contrary to centric fusion, in dissociation one metacentric chromosome with long arms and the second metacentric chromosome with short arms after reciprocal translocation give rise to two acrocentric chromosomes [Fig. 22.11 (b)].
Tandem fusions:
This results from a single break in the vicinity of centromere of one chromosome and another break near the end of a second chromosome. If the two chromosomes involved in this process are acrocentrics, then one large acrocentric and one small metacentric chromosomes will result (Figs. 22.11 (c), 22.12).
But if the interchange takes place between one metacentric and the other acrocentric chromosome, then two acrocentrics, one short and the other large may be formed [Fig. 22.11 (d)].
Translocations are usually non-lethal in their effects. Like deficiency and duplication, the translocations may also be homozygous or heterozygous provided the aberrations in question are not associated with lethality.
The homozygous translocations are characterised by the presence of same gene sequences in the transposed segments of homologous chromosomes. If an organism carrying two pairs of chromosomes with reciprocal translocation is crossed with a normal organism, the offspring will be heterozygous for translocation. Translocation heterozygotes thus possess translocated and normal chromosomes.
The following are the important effects or evidences of translocations:
1. Cytological effects
2. Genetical effects
1. Cytological Effects of Translocations:
The chromosomes of homozygous translocations generally behave as do the normal ones from which they arise, except that new linkage groups are formed. If they persist they can give rise to new chromosomal races in the population.
In the individuals heterozygous for a symmetrical reciprocal translocation (structural hybrids) two chromosomes with translocation and two normal chromosomes share a partial homology, but no two are identical.
Since synapsis is a matter of homologous regions (gene to gene pairing) and these regions are distributed over four chromosomes, in simple reciprocal translocation heterozygote, the association of all four chromosomes will be formed. Such an association will result in a cross configuration (+) at pachytene i.e., there will develop a group of four associated chromosomes (2 normal + 2 translocated) (Fig. 22.14).
Heterozygosity for translocation reduces the crossing over frequency. The crossing over can take place in any of the four pairing segments of cross-like configuration in reciprocal translocation heterozygote but the results will vary according to the cross over site relative to centromeres and breakpoints of the translocation.
If the crossing over occurs in the regions between the centromeres and break points (i.e., interstitial region) it will result in duplicated and deficient chromosomes irrespective of adjacent or alternate distribution patterns.
This may be major cause of sterility in translocation heterozygotes. But the crossing over appears to be restricted in the interstitial regions because of inefficient synapsis between the chromosomes. If the crossing over takes place outside the interstitial regions, it does not affect the segregation patterns since one homologous section is exchanged for another.
Reduced crossing over within the translocated part is most pronounced in the vicinity of the interchange points. It is so on account of non-homologous pairing in interchange region or due to difficulty in its meiotic chromosome pairing.
The subsequent behaviour of this cross configuration depends upon the frequency and locatior of the chiasmata and the centromere orientation. If chiasma formation takes place in all four pairing segments, a ring of four chromosomes results. The occurrence of quadrivalent rings has been observed during metaphase in Datura, peas, wheat, Tradescantia and some other plants as well as animals.
If chiasma formation fails in one of the four pairing segments, a chain of four chromosomes results. If chiasma formation takes place in two adjacent or alternate pairing segments, a chain of three chromosomes and one univalent, or two bivalents will result.
Rings and chains contain structurally normal and structurally changed chromosomes in alternating sequence. Now the distribution of the four chromosomes in ring or chain configuration at anaphase I of meiosis is determined by the orientation of centromeres. There are two common patterns of distribution, adjacent and alternate (Fig. 22.14).
(a) Adjacent distribution:
In this the chromosomes located alternately in the pairing configuration are segregated in such a way that one structurally normal and one translocated chromosome go to one pole and their counterparts, to the opposite pole. In this case both meiotic products are duplicated.
In adjacent distribution there are two events:
(i) Adjacent 1 distribution in which the centromeres of neighbouring non-homologous chromosomes segregate to same pole, and
(ii) Adjacent-2 distribution in which homologous centromeres migrate to the same pole, but it is very rare in occurrence.
(b) Alternate distribution:
In this the two normal chromosomes go to one pole and the two translocated chromosomes go to the opposite pole during anaphase I and so the gametes- formed are of two kinds; some with normal chromosomes and some with translocated chromosomes.
Multiple Translocation System:
If the arm of one of the two translocated chromosomes is involved in a second interchange with a third non-homologous chromosome, a ring of 6 chromosomes (Fig. 22.15) may be formed at metaphase I. In the same way, a third interchange would give rise to a ring of 8 chromosomes.
This process can go on until the entire complement of chromosomes is involved. This produces a translocation complex or complex heterozygote.
In Rhoeo spathacea all 12 chromosomes are involved in reciprocal translocations and as a result of this a ring of 12 chromosomes is formed during meiosis. Oenothera exhibits a similar tendency which varies with the species.
In O. hookeri normally 7 bivalents are formed but in other species formation of multivalent ring and bivalents may be observed and in some other species, such as O. lamarkiana all fourteen chromosomes are linked to form a ring at metaphase I. Translocation complexes of the types described in Rhoeo and Oenothera are not known in animals.
Similar situation has been reported in several other plants exposed to X-rays, gamma-rays and mutagenic chemicals.
2. Genetic Effects of Tanslocations:
The main genetic effects of translocations are as follows:
(i) It brings about a qualitative change in the chromosomes structure or linkage group,
(ii) It brings about change in the sequences of genes in chromosomes which may eventually produce several abnormalities in body characters. This is position effect.
(iii) Semi-sterility. The translocation heterozygotes are generally semi-sterile because they produce gametes containing duplicated and deficient chromosomes as a result of typical pairing behaviours, crossing over and segregation patterns of chromosomes.
The translocation is of great importance for the individuals and the species. The most harmful effect of a reciprocal translocation is the semi-sterility it causes. It also causes severe modifications of the normal developmental pattern.
In the evening primrose (Oenothera) a number of variations are associated with translocations. This was the plant whose variability led De Vries to propose his popular mutation theory.
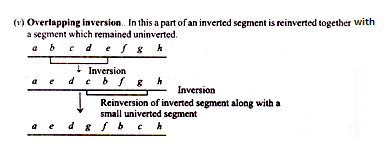
Chromosomal Aberration: Type # 4. Inversion:
I”, is an intrachromosomal aberration characterised by inversion or reversal of a chromosome segment and the gene sequence contained therein relative to the standard chromosome or linkage group in question. It occurs in intercalary segment of the chromosome. There is no experimental evidence for occurrence of inversions in terminal segments of chromosomes.
According to the breakage reunion —model, the intercalary inversions are formed when two breaks occur in a chromosome, the middle segment between the break points (referred to as inversion points) is inverted or rotated through : and then reunion of the three segments at the sites of breakage takes place as shown below Fig. 22.16).
Sometimes inverted segment or a part of it may again undergo inversion. This is called included inversion. Thus, in the above example, if the segment, GFEDC in the inverted chromosome undergoes further inversion, the result will be that the chromosome will regain original gene sequence ABCDEFGHIJ.
Types of Inversions:
The inversions are classified according to the number of inverted segments within the chromosome and the location of inversion points with respect to each other.
1. Single inversions:
When a chromosome contains a single inverted segment, it is called single inversion. Single inversions are classified according to whether or not the inverted segment of the chromosome carries the centromere.
(i) Paracentric inversion. This is the most common type of inversion which is confined to a single arm of a chromosome. In this, the inverted segment of the chromosome does not carry centromere. It is also called acentric or dyscentric orparakinetic or asymmetrical inversion (Fig. 22.17).
(ii) Pericentric inversion:
In this type of inversion, the break points are located in both the arms of chromosome so that the inverted segment includes centromere (Fig. 22.17). It is also called transcentric or eucentric or transkinetic or symmetrical inversion. If the two breakpoints involved in the formation of a pericentric inversion are equidistant from the centromere, the inverted chromosome will appear morphologically similar to normal one.
But, if the breakpoints are asymmetric (not equidistant) from the centromere, a shift of centromere from acrocentric to metacentric or vice-versa may occur, thus causing a marked change in the appearance of chromosome. This indicates that pericentric inversions might have played important role in the evolution of new karyotypes.
2. Complex inversions:
When a chromosome contains more than one inverted segment, it is called complex inversion.
The complex inversions are classified as follows:
(i) Independent inversion:
When the inverted segments are separated from one another by uninverted segment, the inversion is said to be independent inversion.
Inversions may be either homozygous or heterozygous. Homozygous inversions have homologous chromosomes with identical inversions. They show normal meiotic pairing and distribution. Heterozygous inversions have one homologue with inversion and the other homologue without inversion (i.e., normal).
Inversion heterozygotes show important cytological and genetic effects. Because there is no net loss or gain of genetic material, inversion heterozygotes are perfectly viable. The pairing behaviour of inversion chromosome with standard or non-inverted homologue depends on the length of inversion and the longitudinal relationship of the inverted and uninverted chromosome segments.
If the inversion is long, chromosome pairing involves formation of characteristic loop in the normal homologous chromosome and the inversion chromosome pairs with the normal chromosome in such a way that homologous loci pair with each other. The location of the inverted segment, thus, can be recognised cytologically by the presence of an inversion loop in the paired homologues during meiosis.
If the inverted segment is so small that loop formation is not possible either the inverted segment is left unpaired or it may pair with non-homologous segment of normal chromosome. The size of the loop is a function of the size of the inversion—the larger is the inversion, the larger will be the loop.
The crossing over and chiasma formation within and outside inversion loop give rise to secondary structural changes (duplication and deletion) depending upon the type of inversion (paracentric or pericentric), the number of chiasmata and localization of chiasmata. Such structural changes result in meiotic products with unbalanced sets of chromosomes.
Single crossing over within a pericentric heterozygous inversion produces two normal meiotic products and two abnormal products—containing chromosom that are either duplicated or deficient for certain gene loci a (Fig. 22.18 B).
In plants gametes containing duplication or deficiency are generally not viable. So, pericentric inversion heterozygotes are semi-sterile, although more than 50% viable. In animals, the gametes with duplication and deficiency in chromosomes are usually normal in function, but zygote usually does not survive.
Single crossing over within a paracentric inversion has more complex consequences and it produces one chromosome with two centromere (dicentric chromosome) and one with no centromere (acentric chromosome segment).
During anaphase 1 of meiosis dicentric chromosome is pulled as a bridge between two spindle poles and acentric chromosome, because it has no spindle fibre attachment, floats randomly as laggard and is eventually lost (Fig. 22.18 A).
The dicentric bridge may break at any point giving rise to duplications and deficiencies in the meiotic products. Remarkably, in Drosophila and plant egg cells, the dicentric bridge may remain intact long after anaphase I. Thus, the two daughter nuclei either will be linked by dicentric bridge or will contain the fragments of the bridge if it breaks.
The fragment associated with the bridge has the effect of a deficiency and the size of deficiency determines the reduction in fertility. That is, a meiotic product or gamete lacking in one or more genes because of the loss of a segment of chromosome is likely to be non-viable and hence sterile.
In general, females heterozygous for paracentric inversion manifest no serious sterility because in most cases the chromosomes are oriented in specific direction during gametogenesis which facilitates exclusion of dicentric and acentric chromosomes from functional gametes.
When two cross overs are formed within inversion loop, the result will depend upon the number of chromatids involved. Two strand-double crossing over will yield four normal chromatids, two of which will be involved in crossing over and the other will were not. Such a condition can be detected only when appropriate genetic markers are present within the region of crossing over.
Three-strand double crossing over will yield one non-cross over chromatid, one cross-over chromatid and two acentric fragments. Four-strand double cross-over would yield two dicentric chromatids and two acentric fragments and deficiency and duplication would presumably lead to in-viability of the meiotic products or gametes or would cause death of zygote or embryo if such gametes were involved in fertilization.
It is, therefore, evident that paracentric inversions have a drastic effect on the recovery of chromatids involved in crossing over.
Another major effect of inversions is to suppress the recombination of genie loci by crossing over to maintain in the population a specific segment of a chromosome.
Crossing over does occur within the inversion, but the crossing over products do not usually contribute to the next generation either because the gametes or zygotes are inviable or because the cross over chromosomes are eliminated from non-functional megaspore in plants or polar bodies in the animals.
The presence of recessive lethal gene within inverted segment can be of added advantage in preserving structural heterozygosity because heterozygotes for lethal recessive genes will be viable and homozygous, nonviable.
In CIB stock of Drosophila, C factor which is a cross-over suppressor is found to be inverted segment of chromosome, 1 component, a recessive lethal, prevents homozygosity for CIB chromosome and B gene accounts for bar eye.
In CIB chromosome C factor is flanked on either side by two marker genes 1 and B. Muller (1928) was the first to take advantage of cross-over suppressing property o inversion heterozygote to detect sex-linked recessive lethal mutations in Drosophila induced by X rays.
ADVERTISEMENTS:
The genetic evidence of inversion thus will be:
(i) Suppression of crossing over, and
(ii) Possibly; the appearance of mutation owing to position effect.
As mentioned earlier, inversion and translocation involve no loss or gain in the genes as such. Simply they bring about change in the position of some genes and no gene mutation (i.e., no change in the nature of gene) is involved in these cases.
The changed positions of genes in the chromosomes may have important consequences since continuous genes sometimes are concerned in the completion of related steps of some repetition biochemical reactions. All such alterations of gene functions due to change in the sequence of genes are referred to as “position effects”.
Inversion homozygotes can be detected cytologically and genetically in the following ways:
(i) By detecting changed linkage relation with the help of genetic linkage studies,
(ii) By detecting the changes in chromosome morphology during mitotic metaphase.
(iii) By observing the changes in chromosome bands.
The inversion heterozygotes are detected by the following characteristics:
(i) Formation of inversion loop during the prophase I of meiosis.
(ii) Formation of dicentric chromosomal bridge, acentric fragments during anaphase I.
(iii) Development of abnormal meiotic products which may be detected by means of tetrad analysis.
(iv) Decreased fertility resulting due to production of genetically unbalanced meiotic products or gametes via crossing over.
(v) Reduced crossing-over frequency